Add analysis
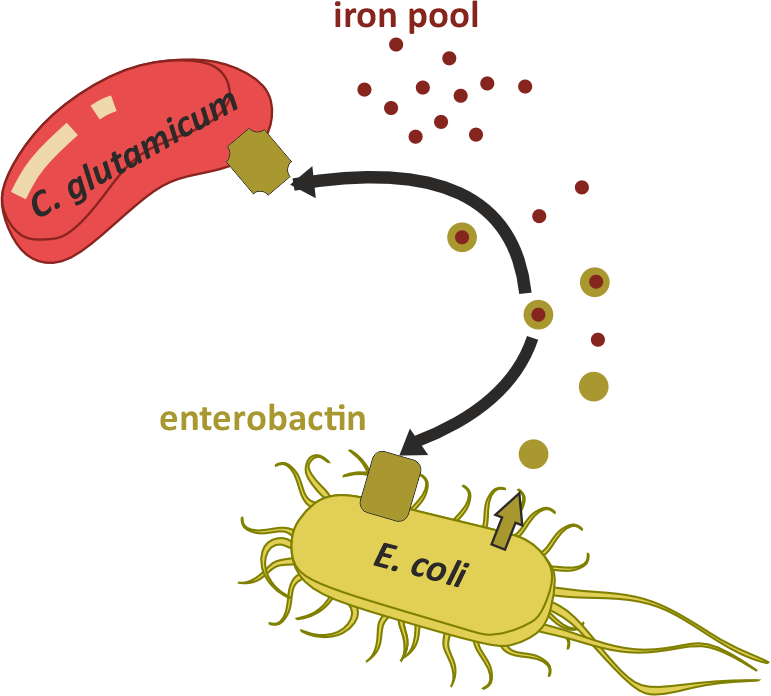
Dynamic Entrobaction model
tune
Simulation parameters
analytics
Analyses
Time course
Loading chart...
Innoculation ratio scan
Model Details
Review and edit model structure, biological variables, and kinetic parameters.
| Name | Tex name | Initial value | Actions |
|---|---|---|---|
Edit Options
Edit Options
Edit Options
Edit Options
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Leaves
Arithmetic
Functions
Trigonometry
Hyperbolic
Comparison
Logic
Equation builder
innoculationRatio
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Preview
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Leaves
Arithmetic
Functions
Trigonometry
Hyperbolic
Comparison
Logic
Equation builder
1
innoculationRatio
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Preview
preview
Generated Python Code
import numpy as np
def model(
time: float,
variables: list[float],
):
x1, x2, s1, p1 = variables
Y_X1_S = 0.45
Y_X2_S = 0.5
mu_max1 = 0.22
mu_max2 = 0.45
K_s1 = 0.0005
K_s2 = 0.005
q_p1_max = 0.015
q_up_X1_max = 0.005
q_up_X2_max = 0.01
K_s_X1_minus_P = 0.00001
K_s_X2_minus_P = 0.001
innoculationRatio = 0.5
mu1 = (mu_max1 * p1 * s1) / ((K_s1 + s1) * (K_s_X1_minus_P + p1))
mu2 = (mu_max2 * p1 * s1) / ((K_s2 + s1) * (K_s_X2_minus_P + p1))
q_p1 = (mu1 * q_p1_max) / (mu_max1)
q_up1 = (mu1 * q_up_X1_max) / (mu_max1)
q_up2 = (mu2 * q_up_X2_max) / (mu_max2)
dx1dt = +(x1)*mu1
dx2dt = +(x2)*mu2
ds1dt = +(- (x1) / (Y_X1_S))*mu1+(- (x2) / (Y_X2_S))*mu2
dp1dt = +(x1)*q_p1+(- x1)*q_up1+(- x2)*q_up2
return [dx1dt, dx2dt, ds1dt, dp1dt]
def all_derived(
time: float,
variables: list[float],
):
x1, x2, s1, p1 = variables
Y_X1_S = 0.45
Y_X2_S = 0.5
mu_max1 = 0.22
mu_max2 = 0.45
K_s1 = 0.0005
K_s2 = 0.005
q_p1_max = 0.015
q_up_X1_max = 0.005
q_up_X2_max = 0.01
K_s_X1_minus_P = 0.00001
K_s_X2_minus_P = 0.001
innoculationRatio = 0.5
mu1 = (mu_max1 * p1 * s1) / ((K_s1 + s1) * (K_s_X1_minus_P + p1))
mu2 = (mu_max2 * p1 * s1) / ((K_s2 + s1) * (K_s_X2_minus_P + p1))
q_p1 = (mu1 * q_p1_max) / (mu_max1)
q_up1 = (mu1 * q_up_X1_max) / (mu_max1)
q_up2 = (mu2 * q_up_X2_max) / (mu_max2)
return [mu1, mu2, q_p1, q_up1, q_up2]
derived = all_derived
y0 = {"x1": 0.5, "x2": 0.5, "s1": 10, "p1": 0.02}
preview
Generated LaTeX Code
\begin{align*}
\frac{d x1}{dt} &= x1 \cdot \frac{mu\_max1 \cdot p1 \cdot s1}{(K\_s1 + s1) \cdot (K\_s\_X1-P + p1)}\\
\frac{d x2}{dt} &= x2 \cdot \frac{mu\_max2 \cdot p1 \cdot s1}{(K\_s2 + s1) \cdot (K\_s\_X2-P + p1)}\\
\frac{d s1}{dt} &= - \frac{x1}{Y\_X1\_S} \cdot \frac{mu\_max1 \cdot p1 \cdot s1}{(K\_s1 + s1) \cdot (K\_s\_X1-P + p1)} \\
& - \frac{x2}{Y\_X2\_S} \cdot \frac{mu\_max2 \cdot p1 \cdot s1}{(K\_s2 + s1) \cdot (K\_s\_X2-P + p1)}\\
\frac{d p1}{dt} &= x1 \cdot \frac{mu1 \cdot q\_p1\_max}{mu\_max1} \\
& - x1 \cdot \frac{mu1 \cdot q\_up\_X1\_max}{mu\_max1} \\
& - x2 \cdot \frac{mu2 \cdot q\_up\_X2\_max}{mu\_max2}
\end{align*}Edit analysis
Plot options
Variable selection
Show Normalize
Edit parameter scan
Plot options
Variable selection
Show Normalize