Add analysis
Ebeling 2026 model
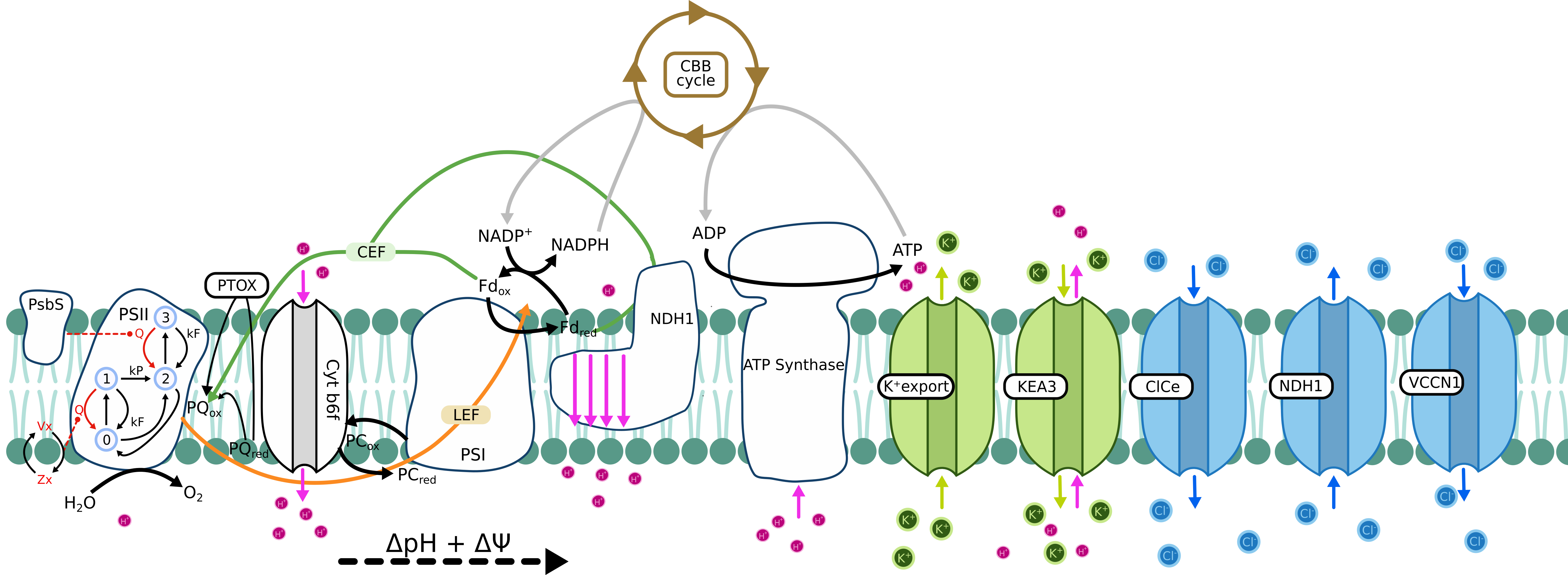
Simulation parameters
Analyses
ECS
Loading chart...
PAM Fluorescence
Loading chart...
Model Details
Review and edit model structure, biological variables, and kinetic parameters.
| Name | Tex name | Initial value | Actions |
|---|---|---|---|
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Generated Python Code
import numpy as np
def model(
time: float,
variables: list[float],
):
_3PGA, BPGA, GAP, DHAP, FBP, F6P, G6P, G1P, SBP, S7P, E4P, X5P, R5P, RUBP, RU5P, ATP, Ferredoxine_oxidised, Light_minus_harvesting_complex, NADPH, Plastocyanine_oxidised, Plastoquinone_oxidised, PsbS_de_minus_protonated, Violaxanthin, MDA, H2O2, DHA, GSSG, Thioredoxin_oxidised, E_inactive, P700FA, P700_plus_FA_minus, P700FA_minus, B0, B1, B2, pH_lumen, pH, ATPactivity, delta_psi, K_stroma, Cl_stroma = variables
PPFD = 100
CO2_dissolved = 0.013226
O2_dissolved_lumen = 8
bH = 100
F = 96.485
E0_PC = 0.38
E0_P700 = 0.48
E0_FA = -0.55
E0_Fd = -0.43
E0_NADP = -0.113
convf = 0.032
R = 0.0083
T = 298
Carotenoids_tot = 1
Fd_star = 5
PC_tot = 4
PSBS_tot = 1
LHC_tot = 1
gamma0 = 0.06260060801266355
gamma1 = 0.4053583123566203
gamma2 = 0.7040758738825375
gamma3 = 0.07834807781016208
kZSat = 0.12
E0_QA = -0.14
E0_PQ = 0.354
PQ_tot = 17.5
staticAntII = 0.1
staticAntI = 0.37
Thioredoxin_tot = 1
E_total = 6
NADP_star = 0.8
A_star_P = 2.55
Pi_tot = 17.05
kf_ferredoxin_thioredoxin_reductase = 0.8
kf_tr_activation = 1
kf_tr_inactivation = 0.1
ASC_tot_star = 10
Glutathion_tot = 10
kf_atp_synthase = 20
HPR = 4.666666666666667
Pi_mol = 0.01
DeltaG0_ATP = 30.6
kh_lhc_protonation = 10
kf_lhc_protonation = 0.15837051384170664
ksat_lhc_protonation = 6.2539066418842255
kf_lhc_deprotonation = 0.015892570403695704
kf_cyclic_electron_flow = 1
kf_violaxanthin_deepoxidase = 0.0006091912188339879
kh_violaxanthin_deepoxidase = 4
ksat_violaxanthin_deepoxidase = 6.193595407850397
kf_zeaxanthin_epoxidase = 0.000106261953934132
km_fnr_Ferredoxine_reduced = 1.56
km_fnr_NADP = 0.22
E0_fnr = 3
kcat_fnr = 500
kf_ndh = 0.002
PSII_total = 2.5
PSI_total = 2.5
kH0 = 500000000
kPQred = 250
kPCox = 2500
kFdred = 250000
k2 = 5000000000
kH = 5000000000
kF = 625000000
kMehler = 1
kf_proton_leak = 10
kPTOX = 0.01
kStt7 = 0.0035
km_lhc_state_transition_12 = 0.2
n_ST = 2
kPph1 = 0.0013
E0_rubisco = 1
kcat_rubisco_carboxylase = 2.72
km_rubisco_carboxylase_RUBP = 0.02
km_rubisco_carboxylase_CO2_dissolved = 0.0107
ki_rubisco_carboxylase_3PGA = 0.04
ki_rubisco_carboxylase_FBP = 0.04
ki_rubisco_carboxylase_SBP = 0.075
ki_rubisco_carboxylase_Orthophosphate = 0.9
ki_rubisco_carboxylase_NADPH = 0.07
kre_phosphoglycerate_kinase = 800000000
keq_phosphoglycerate_kinase = 0.00031
kre_gadph = 800000000
keq_gadph = 16000000
kre_triose_phosphate_isomerase = 800000000
keq_triose_phosphate_isomerase = 22
kre_aldolase_dhap_gap = 800000000
keq_aldolase_dhap_gap = 7.1
kre_aldolase_dhap_e4p = 800000000
keq_aldolase_dhap_e4p = 13
E0_fbpase = 1
kcat_fbpase = 1.6
km_fbpase_s = 0.03
ki_fbpase_F6P = 0.7
ki_fbpase_Orthophosphate = 12
kre_transketolase_gap_f6p = 800000000
keq_transketolase_gap_f6p = 0.084
kre_transketolase_gap_s7p = 800000000
keq_transketolase_gap_s7p = 0.85
E0_SBPase = 1
kcat_SBPase = 0.32
km_SBPase_s = 0.013
ki_SBPase_Orthophosphate = 12
kre_ribose_phosphate_isomerase = 800000000
keq_ribose_phosphate_isomerase = 0.4
kre_ribulose_phosphate_epimerase = 800000000
keq_ribulose_phosphate_epimerase = 0.67
E0_phosphoribulokinase = 1
kcat_phosphoribulokinase = 7.9992
km_phosphoribulokinase_RU5P = 0.05
km_phosphoribulokinase_ATP = 0.05
ki_phosphoribulokinase_3PGA = 2
ki_phosphoribulokinase_RUBP = 0.7
ki_phosphoribulokinase_Orthophosphate = 4
ki_phosphoribulokinase_4 = 2.5
ki_phosphoribulokinase_5 = 0.4
kre_g6pi = 800000000
keq_g6pi = 2.3
kre_phosphoglucomutase = 800000000
keq_phosphoglucomutase = 0.058
Orthophosphate_external = 0.5
km_ex_pga = 0.25
km_ex_gap = 0.075
km_ex_dhap = 0.077
km_N_translocator_Orthophosphate_external = 0.74
km_N_translocator_Orthophosphate = 0.63
kcat_N_translocator = 2
E0_N_translocator = 1
E0_ex_g1p = 1
km_ex_g1p_G1P = 0.08
km_ex_g1p_ATP = 0.08
ki_ex_g1p = 10
ki_ex_g1p_3PGA = 0.1
ki_ex_g1p_F6P = 0.02
ki_ex_g1p_FBP = 0.02
kcat_ex_g1p = 0.32
kf_mda_reductase_1 = 500
E0_mda_reductase_2 = 0.002
kcat_mda_reductase_2 = 300
km_mda_reductase_2_NADPH = 0.023
km_mda_reductase_2_MDA = 0.0014
kf1 = 10000
kr1 = 220
kf2 = 10000
kr2 = 4000
kf3 = 2510
kf4 = 10000
kr4 = 4000
kf5 = 2510
XT = 0.07
E0_glutathion_reductase = 0.0014
kcat_glutathion_reductase = 595
km_glutathion_reductase_NADPH = 0.003
km_glutathion_reductase_GSSG = 0.2
km_dehydroascorbate_reductase_DHA = 0.07
km_dehydroascorbate_reductase_GSH = 2.5
K = 0.5
E0_dehydroascorbate_reductase = 0.0017
kcat_dehydroascorbate_reductase = 142
kf_ex_atp = 0.5
kf_ex_nadph = 0.5
kH_Qslope = 30000000000
b6f_content = 1
max_b6f = 500
pKreg = 7
stroma_buffering = 400
kActATPase = 0.001
kDeactATPase = 0.002
k_ATPsynthase = 20
b = 1.8688304401249531
pK0E = 5.960025833706074
k_import_ATP = 0.5
k_import_NADPH = 0.5
volts_per_charge = 0.000769481926574965
ClCe_PQ = 15.87880046767565
Cl_leak_PQ = 14.92901445507139
KEA3_ATP_treshold = 0.26274793681796166
KEA3_pH_reg = 7.69
K_delta_psi_treshold = 0.08146807307624158
VCCN_delta_psi_treshold = 0.08000900979332677
k_Cl_leak = 25
k_NDH1 = 7.447430768265866
k_KEA = 90
perm_K = 1.6113135416150155
k_VCCN1 = 0.5
k_ClCe = 0.5
K_total = 60
Cl_total = 50
ClCe_ATP_threshold = 0.2
RT = R * T
dG_pH = 2.302585092994046 * R * T
Zeaxanthin = Carotenoids_tot + - Violaxanthin
Ferredoxine_reduced = Fd_star + - Ferredoxine_oxidised
Plastocyanine_reduced = PC_tot + - Plastocyanine_oxidised
PsbS_protonated = PSBS_tot + - PsbS_de_minus_protonated
Light_minus_harvesting_complex_protonated = LHC_tot + - Light_minus_harvesting_complex
Q = PsbS_de_minus_protonated * Violaxanthin * gamma0 + PsbS_protonated * Violaxanthin * gamma1 + (PsbS_de_minus_protonated * Zeaxanthin * gamma3) / (Zeaxanthin + kZSat) + (PsbS_protonated * Zeaxanthin * gamma2) / (Zeaxanthin + kZSat)
keq_Plastoquinone_reduced = np.exp((2 * E0_PQ * F + - 2 * E0_QA * F + - 2 * dG_pH * pH) / (RT))
Plastoquinone_reduced = PQ_tot + - Plastoquinone_oxidised
PSII_cross_section = staticAntII + Light_minus_harvesting_complex * (1 + - staticAntI + - staticAntII)
Thioredoxin_reduced = Thioredoxin_tot + - Thioredoxin_oxidised
E_active = E_total + - E_inactive
NADP = NADP_star + - NADPH
ADP = A_star_P + - ATP
Orthophosphate = Pi_tot + - ATP + - DHAP + - E4P + - F6P + - G1P + - G6P + - GAP + - R5P + - RU5P + - S7P + - X5P + - _3PGA + - 2 * BPGA + - 2 * FBP + - 2 * RUBP + - 2 * SBP
ascorbate = ASC_tot_star + - DHA + - MDA
GSH = Glutathion_tot + - 2 * GSSG
keq_atp_synthase = Pi_mol * np.exp((- DeltaG0_ATP + HPR * dG_pH * (pH + - pH_lumen)) / (RT))
keq_fnr = np.exp((- dG_pH * pH + 2 * E0_NADP * F + - 2 * E0_Fd * F) / (RT))
vmax_fnr = E0_fnr * kcat_fnr
E0_rubisco_active = E0_rubisco * E_active
vmax_rubisco_carboxylase = E0_rubisco_active * kcat_rubisco_carboxylase
E0_fbpase_active = E0_fbpase * E_active
vmax_fbpase = E0_fbpase_active * kcat_fbpase
E0_SBPase_active = E0_SBPase * E_active
vmax_SBPase = E0_SBPase_active * kcat_SBPase
E0_phosphoribulokinase_active = E0_phosphoribulokinase * E_active
vmax_phosphoribulokinase = E0_phosphoribulokinase_active * kcat_phosphoribulokinase
vmax_ex_pga = E0_N_translocator * kcat_N_translocator
N_translocator = 1 + (1 + (km_N_translocator_Orthophosphate_external) / (Orthophosphate_external)) * ((DHAP) / (km_ex_dhap) + (GAP) / (km_ex_gap) + (Orthophosphate) / (km_N_translocator_Orthophosphate) + (_3PGA) / (km_ex_pga))
E0_ex_g1p_active = E0_ex_g1p * E_active
vmax_ex_g1p = E0_ex_g1p_active * kcat_ex_g1p
vmax_mda_reductase_2 = E0_mda_reductase_2 * kcat_mda_reductase_2
vmax_glutathion_reductase = E0_glutathion_reductase * kcat_glutathion_reductase
vmax_dehydroascorbate_reductase = E0_dehydroascorbate_reductase * kcat_dehydroascorbate_reductase
keq_PCP700 = np.exp((E0_P700 * F + - E0_PC * F) / (RT))
keq_FAFd = np.exp((E0_Fd * F + - E0_FA * F) / (RT))
B3 = PSII_total + - B0 + - B1 + - B2
P700_plus_FA = PSI_total + - P700FA + - P700FA_minus + - P700_plus_FA_minus
rel_P700_plus_FA = (P700_plus_FA) / (PSI_total)
rel_P700FA = (P700FA) / (PSI_total)
rel_P700FA_minus = (P700FA_minus) / (PSI_total)
rel_P700_plus_FA_minus = (P700_plus_FA_minus) / (PSI_total)
rel_P700 = (P700_plus_FA + P700_plus_FA_minus) / (PSI_total)
rel_P700_plus = (P700_plus_FA + P700_plus_FA_minus) / (PSI_total)
rel_B0 = (B0) / (PSII_total)
rel_B1 = (B1) / (PSII_total)
rel_B2 = (B2) / (PSII_total)
rel_B3 = (B3) / (PSII_total)
Fluo = (B0 * PSII_cross_section * kF) / (k2 + kF + kH0 + Q * kH_Qslope) + (B2 * PSII_cross_section * kF) / (kF + kH0 + Q * kH_Qslope)
keq_b6f_dyn = b6f_content * max_b6f * (1 + - (1) / (1 + (10) ** (pH_lumen + - pKreg)))
protons_lumen = 4000 * (10) ** (- pH_lumen)
protons = 31250 * (10) ** (- pH)
ATP_pmf_activity = ((10) ** (- pK0E) * np.exp((F * b * (delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F))) / (RT))) / (1 + (10) ** (- pK0E) * np.exp((F * b * (delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F))) / (RT)))
deltapH = dG_pH * (pH + - pH_lumen)
deltapH_in_volts = (2.302585092994046 * R * T * (pH + - pH_lumen)) / (F)
pmf = deltapH + F * delta_psi
pmf_in_V = delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F)
keq_b6f = np.exp((- 2 * pmf_in_V + 2 * E0_PC * F + 2 * dG_pH * pH_lumen + - 2 * E0_PQ * F) / (RT))
K_lumen = K_total + - K_stroma
Cl_lumen = Cl_total + - Cl_stroma
total_Cl_2 = (Cl_total) ** (2)
total_K_2 = (K_total) ** (2)
KEA3_reg = (0.81) / ((1 + np.exp(100 * KEA3_ATP_treshold + - 100 * ATP)) * (1 + np.exp(1000 * pH + - 1000 * KEA3_pH_reg)))
dG_K_ions = F * (delta_psi + - (RT * np.log((K_stroma) / (K_lumen))) / (F * np.log(10)))
Cl_driving_force = F * (delta_psi + (RT * np.log((Cl_stroma) / (Cl_lumen))) / (F * np.log(10)))
Keq_NDH1 = np.exp((- 4 * pmf + 2 * E0_PQ * F + - 2 * E0_Fd * F + - 2 * dG_pH * pH) / (RT))
ClCe_activation = (0.9) / (1 + np.exp(100 * ATP + - 100 * ClCe_ATP_threshold))
PQ_ox_div_tot = (Plastoquinone_reduced) / (PQ_tot)
Fd_ox_div_tot = (Ferredoxine_reduced) / (Fd_star)
PC_ox_div_tot = (Plastocyanine_reduced) / (PC_tot)
NADPH_div_tot = (NADPH) / (NADP_star)
ATP_div_tot = (ATP) / (A_star_P)
ferredoxin_thioredoxin_reductase = Ferredoxine_reduced * Thioredoxin_oxidised * kf_ferredoxin_thioredoxin_reductase
tr_activation = E_inactive * Thioredoxin_reduced * kf_tr_activation
tr_inactivation = E_active * kf_tr_inactivation
atp_synthase = ATP_pmf_activity * ATPactivity * k_ATPsynthase * ((ADP) / (convf) + - (ATP) / (convf * kf_atp_synthase))
lhc_protonation = (PsbS_de_minus_protonated * kf_lhc_protonation * (protons_lumen) ** (kh_lhc_protonation)) / ((protons_lumen) ** (kh_lhc_protonation) + (4000 * (10) ** (- ksat_lhc_protonation)) ** (kh_lhc_protonation))
lhc_deprotonation = PsbS_protonated * kf_lhc_deprotonation
cyclic_electron_flow = Plastoquinone_oxidised * kf_cyclic_electron_flow * (Ferredoxine_reduced) ** (2)
violaxanthin_deepoxidase = (Violaxanthin * kf_violaxanthin_deepoxidase * (protons_lumen) ** (kh_violaxanthin_deepoxidase)) / ((protons_lumen) ** (kh_violaxanthin_deepoxidase) + (4000 * (10) ** (- ksat_violaxanthin_deepoxidase)) ** (kh_violaxanthin_deepoxidase))
zeaxanthin_epoxidase = Zeaxanthin * kf_zeaxanthin_epoxidase
fnr = (vmax_fnr * ((NADP * ((Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) ** (2)) / (convf * km_fnr_NADP) + - (NADPH * ((Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)) ** (2)) / (convf * keq_fnr * km_fnr_NADP))) / (-1 + (1 + (NADP) / (convf * km_fnr_NADP)) * (1 + ((Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) ** (2) + (Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) + (1 + (NADPH) / (convf * km_fnr_NADP)) * (1 + ((Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)) ** (2) + (Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)))
ndh = Plastoquinone_oxidised * kf_ndh
proton_leak = kf_proton_leak * (protons_lumen + - 4000 * (10) ** (- pH))
PTOX = O2_dissolved_lumen * Plastoquinone_reduced * kPTOX
lhc_state_transition_12 = (1 * Light_minus_harvesting_complex * kStt7) / (1 + ((Plastoquinone_oxidised) / (PQ_tot * km_lhc_state_transition_12)) ** (n_ST))
lhc_state_transition_21 = Light_minus_harvesting_complex_protonated * kPph1
rubisco_carboxylase = (CO2_dissolved * RUBP * vmax_rubisco_carboxylase) / ((CO2_dissolved + km_rubisco_carboxylase_CO2_dissolved) * (RUBP + km_rubisco_carboxylase_RUBP * (1 + (FBP) / (ki_rubisco_carboxylase_FBP) + (NADPH) / (ki_rubisco_carboxylase_NADPH) + (Orthophosphate) / (ki_rubisco_carboxylase_Orthophosphate) + (SBP) / (ki_rubisco_carboxylase_SBP) + (_3PGA) / (ki_rubisco_carboxylase_3PGA))))
phosphoglycerate_kinase = kre_phosphoglycerate_kinase * (ATP * _3PGA + - (ADP * BPGA) / (keq_phosphoglycerate_kinase))
gadph = kre_gadph * (BPGA * NADPH * protons + - (GAP * NADP * Orthophosphate) / (keq_gadph))
triose_phosphate_isomerase = kre_triose_phosphate_isomerase * (GAP + - (DHAP) / (keq_triose_phosphate_isomerase))
aldolase_dhap_gap = kre_aldolase_dhap_gap * (DHAP * GAP + - (FBP) / (keq_aldolase_dhap_gap))
aldolase_dhap_e4p = kre_aldolase_dhap_e4p * (DHAP * E4P + - (SBP) / (keq_aldolase_dhap_e4p))
fbpase = (FBP * vmax_fbpase) / (FBP + km_fbpase_s * (1 + (F6P) / (ki_fbpase_F6P) + (Orthophosphate) / (ki_fbpase_Orthophosphate)))
transketolase_gap_f6p = kre_transketolase_gap_f6p * (F6P * GAP + - (E4P * X5P) / (keq_transketolase_gap_f6p))
transketolase_gap_s7p = kre_transketolase_gap_s7p * (GAP * S7P + - (R5P * X5P) / (keq_transketolase_gap_s7p))
SBPase = (SBP * vmax_SBPase) / (SBP + km_SBPase_s * (1 + (Orthophosphate) / (ki_SBPase_Orthophosphate)))
ribose_phosphate_isomerase = kre_ribose_phosphate_isomerase * (R5P + - (RU5P) / (keq_ribose_phosphate_isomerase))
ribulose_phosphate_epimerase = kre_ribulose_phosphate_epimerase * (X5P + - (RU5P) / (keq_ribulose_phosphate_epimerase))
phosphoribulokinase = (ATP * RU5P * vmax_phosphoribulokinase) / ((RU5P + km_phosphoribulokinase_RU5P * (1 + (Orthophosphate) / (ki_phosphoribulokinase_Orthophosphate) + (RUBP) / (ki_phosphoribulokinase_RUBP) + (_3PGA) / (ki_phosphoribulokinase_3PGA))) * (ATP * (1 + (ADP) / (ki_phosphoribulokinase_4)) + km_phosphoribulokinase_ATP * (1 + (ADP) / (ki_phosphoribulokinase_5))))
g6pi = kre_g6pi * (F6P + - (G6P) / (keq_g6pi))
phosphoglucomutase = kre_phosphoglucomutase * (G6P + - (G1P) / (keq_phosphoglucomutase))
ex_pga = (_3PGA * vmax_ex_pga) / (N_translocator * km_ex_pga)
ex_gap = (GAP * vmax_ex_pga) / (N_translocator * km_ex_gap)
ex_dhap = (DHAP * vmax_ex_pga) / (N_translocator * km_ex_dhap)
ex_g1p = (ATP * G1P * vmax_ex_g1p) / ((G1P + km_ex_g1p_G1P) * ((1 + (ADP) / (ki_ex_g1p)) * (ATP + km_ex_g1p_ATP) + (Orthophosphate * km_ex_g1p_ATP) / (F6P * ki_ex_g1p_F6P + FBP * ki_ex_g1p_FBP + _3PGA * ki_ex_g1p_3PGA)))
mda_reductase_1 = kf_mda_reductase_1 * (MDA) ** (2)
mda_reductase_2 = (MDA * NADPH * vmax_mda_reductase_2) / (MDA * NADPH + MDA * km_mda_reductase_2_NADPH + NADPH * km_mda_reductase_2_MDA + km_mda_reductase_2_MDA * km_mda_reductase_2_NADPH)
ascorbate_peroxidase = (H2O2 * XT * ascorbate) / ((H2O2) / (kf2) + (H2O2) / (kf4) + (ascorbate) / (kf1) + H2O2 * ascorbate * ((1) / (kf3) + (1) / (kf5)) + (kr1) / (kf1 * kf2) + (H2O2 * kr2) / (kf2 * kf3) + (H2O2 * kr4) / (kf4 * kf5) + (kr1 * kr2) / (kf1 * kf2 * kf3))
glutathion_reductase = (GSSG * NADPH * vmax_glutathion_reductase) / (GSSG * NADPH + GSSG * km_glutathion_reductase_NADPH + NADPH * km_glutathion_reductase_GSSG + km_glutathion_reductase_GSSG * km_glutathion_reductase_NADPH)
dehydroascorbate_reductase = (DHA * GSH * vmax_dehydroascorbate_reductase) / (K + DHA * GSH + DHA * km_dehydroascorbate_reductase_GSH + GSH * km_dehydroascorbate_reductase_DHA)
toP700FA_minus = P700_plus_FA_minus * Plastocyanine_reduced * kPCox + - (P700FA_minus * Plastocyanine_oxidised * kPCox) / (keq_PCP700)
toP700FA_v3 = Ferredoxine_oxidised * P700FA_minus * kFdred + - (Ferredoxine_reduced * P700FA * kFdred) / (keq_FAFd)
toP700_plus_FA = Ferredoxine_oxidised * P700_plus_FA_minus * kFdred + - (Ferredoxine_reduced * P700_plus_FA * kFdred) / (keq_FAFd)
toP700FA_v5 = P700_plus_FA * Plastocyanine_reduced * kPCox + - (P700FA * Plastocyanine_oxidised * kPCox) / (keq_PCP700)
PSI = P700FA * PPFD * (1 + - PSII_cross_section)
mehler1 = O2_dissolved_lumen * P700FA_minus * kMehler
mehler2 = O2_dissolved_lumen * P700_plus_FA_minus * kMehler
B01 = B0 * PPFD * PSII_cross_section
B10Q = B1 * (kH0 + Q * kH_Qslope)
B10F = B1 * kF
B12 = B1 * k2
B20 = B2 * Plastoquinone_oxidised * kPQred + - (B0 * Plastoquinone_reduced * kPQred) / (keq_Plastoquinone_reduced)
B23 = B2 * PPFD * PSII_cross_section
B32F = B3 * kF
B32Q = B3 * (kH0 + Q * kH_Qslope)
b6f = (Plastocyanine_oxidised * Plastoquinone_reduced * keq_b6f_dyn) / (Plastoquinone_oxidised + Plastoquinone_reduced) + - (Plastocyanine_reduced * keq_b6f_dyn * (1 + - (Plastoquinone_reduced) / (Plastoquinone_oxidised + Plastoquinone_reduced))) / (keq_b6f)
vATPactivity = kActATPase * (1 + - ATPactivity) if PPFD > 0 else - ATPactivity * kDeactATPase
vATP_shuttle = ADP * k_import_ATP + - ATP * kf_ex_atp
vNADPH_shuttle = NADP * k_import_NADPH + - NADPH * kf_ex_nadph
KEA3 = max(0, KEA3_reg * k_KEA * (K_stroma * protons_lumen + - K_lumen * protons))
voltage_K_channel = (0.9 * K_lumen * dG_K_ions * perm_K) / (K_stroma * (1 + np.exp(1000 * K_delta_psi_treshold + - 1000 * delta_psi)))
VCCN1 = (0.9 * Cl_driving_force * Cl_stroma * k_VCCN1) / (Cl_lumen * (1 + np.exp(1000 * VCCN_delta_psi_treshold + - 1000 * delta_psi)))
Cl_leak = (0.9 * k_Cl_leak * (Cl_lumen + - Cl_stroma) ** (2)) / (total_Cl_2 * (1 + np.exp(10 * Cl_leak_PQ + - 10 * Plastoquinone_oxidised)))
NDH1 = (0.9 * Plastoquinone_oxidised * k_NDH1 * (10) ** (-6.5 + pH_lumen) * (Ferredoxine_reduced) ** (2)) / ((0.5 + (10) ** (-6.5 + pH_lumen)) * (1 + 7.38905609893065 * np.exp(- 100 * P700_plus_FA_minus)))
ClCe_bi = ClCe_activation * k_ClCe * (Cl_stroma + - Cl_lumen)
d_3PGAdt = +2*rubisco_carboxylase-phosphoglycerate_kinase-ex_pga
dBPGAdt = +phosphoglycerate_kinase-gadph
dGAPdt = +gadph-triose_phosphate_isomerase-aldolase_dhap_gap-transketolase_gap_f6p-transketolase_gap_s7p-ex_gap
dDHAPdt = +triose_phosphate_isomerase-aldolase_dhap_gap-aldolase_dhap_e4p-ex_dhap
dFBPdt = +aldolase_dhap_gap-fbpase
dF6Pdt = +fbpase-transketolase_gap_f6p-g6pi
dG6Pdt = +g6pi-phosphoglucomutase
dG1Pdt = +phosphoglucomutase-ex_g1p
dSBPdt = +aldolase_dhap_e4p-SBPase
dS7Pdt = -transketolase_gap_s7p+SBPase
dE4Pdt = -aldolase_dhap_e4p+transketolase_gap_f6p
dX5Pdt = +transketolase_gap_f6p+transketolase_gap_s7p-ribulose_phosphate_epimerase
dR5Pdt = +transketolase_gap_s7p-ribose_phosphate_isomerase
dRUBPdt = -rubisco_carboxylase+phosphoribulokinase
dRU5Pdt = +ribose_phosphate_isomerase+ribulose_phosphate_epimerase-phosphoribulokinase
dATPdt = +(convf)*atp_synthase-phosphoglycerate_kinase-phosphoribulokinase-ex_g1p+vATP_shuttle
dFerredoxine_oxidiseddt = +ferredoxin_thioredoxin_reductase+2*cyclic_electron_flow+2*fnr-toP700FA_v3-toP700_plus_FA+2*NDH1
dLight_minus_harvesting_complexdt = -lhc_state_transition_12+lhc_state_transition_21
dNADPHdt = +(convf)*fnr-gadph-mda_reductase_2-glutathion_reductase+vNADPH_shuttle
dPlastocyanine_oxidiseddt = +toP700FA_minus+toP700FA_v5-2*b6f
dPlastoquinone_oxidiseddt = -cyclic_electron_flow-ndh+PTOX-0.5*B20+b6f-NDH1
dPsbS_de_minus_protonateddt = -lhc_protonation+lhc_deprotonation
dViolaxanthindt = -violaxanthin_deepoxidase+zeaxanthin_epoxidase
dMDAdt = -2*mda_reductase_1-2*mda_reductase_2+2*ascorbate_peroxidase
dH2O2dt = -ascorbate_peroxidase+(convf)*mehler1+(convf)*mehler2
dDHAdt = +mda_reductase_1-dehydroascorbate_reductase
dGSSGdt = -glutathion_reductase+dehydroascorbate_reductase
dThioredoxin_oxidiseddt = -ferredoxin_thioredoxin_reductase+5*tr_activation
dE_inactivedt = -5*tr_activation+5*tr_inactivation
dP700FAdt = +toP700FA_v3+toP700FA_v5-PSI+2*mehler1
dP700_plus_FA_minusdt = -toP700FA_minus-toP700_plus_FA+PSI-2*mehler2
dP700FA_minusdt = +toP700FA_minus-toP700FA_v3-2*mehler1
dB0dt = -B01+B10Q+B10F+B20
dB1dt = +B01-B10Q-B10F-B12
dB2dt = +B12-B20-B23+B32F+B32Q
dpH_lumendt = +0.04666666666666667*atp_synthase+0.01*proton_leak-0.01*B12-0.04*b6f+0.01*KEA3-0.04*NDH1
dpHdt = -0.011666666666666667*atp_synthase-0.0025*proton_leak+0.0025*B20+0.01*b6f-0.0025*KEA3+0.01*NDH1
dATPactivitydt = +vATPactivity
ddelta_psidt = +(- HPR * volts_per_charge)*atp_synthase+(- 1 * volts_per_charge)*proton_leak+(volts_per_charge)*B12+(4 * volts_per_charge)*b6f+(- 1 * volts_per_charge)*voltage_K_channel+(- 1 * volts_per_charge)*VCCN1+(volts_per_charge)*Cl_leak+(4 * volts_per_charge)*NDH1
dK_stromadt = -KEA3+voltage_K_channel
dCl_stromadt = -VCCN1+Cl_leak-ClCe_bi
return [d_3PGAdt, dBPGAdt, dGAPdt, dDHAPdt, dFBPdt, dF6Pdt, dG6Pdt, dG1Pdt, dSBPdt, dS7Pdt, dE4Pdt, dX5Pdt, dR5Pdt, dRUBPdt, dRU5Pdt, dATPdt, dFerredoxine_oxidiseddt, dLight_minus_harvesting_complexdt, dNADPHdt, dPlastocyanine_oxidiseddt, dPlastoquinone_oxidiseddt, dPsbS_de_minus_protonateddt, dViolaxanthindt, dMDAdt, dH2O2dt, dDHAdt, dGSSGdt, dThioredoxin_oxidiseddt, dE_inactivedt, dP700FAdt, dP700_plus_FA_minusdt, dP700FA_minusdt, dB0dt, dB1dt, dB2dt, dpH_lumendt, dpHdt, dATPactivitydt, ddelta_psidt, dK_stromadt, dCl_stromadt]
def all_derived(
time: float,
variables: list[float],
):
_3PGA, BPGA, GAP, DHAP, FBP, F6P, G6P, G1P, SBP, S7P, E4P, X5P, R5P, RUBP, RU5P, ATP, Ferredoxine_oxidised, Light_minus_harvesting_complex, NADPH, Plastocyanine_oxidised, Plastoquinone_oxidised, PsbS_de_minus_protonated, Violaxanthin, MDA, H2O2, DHA, GSSG, Thioredoxin_oxidised, E_inactive, P700FA, P700_plus_FA_minus, P700FA_minus, B0, B1, B2, pH_lumen, pH, ATPactivity, delta_psi, K_stroma, Cl_stroma = variables
PPFD = 100
CO2_dissolved = 0.013226
O2_dissolved_lumen = 8
bH = 100
F = 96.485
E0_PC = 0.38
E0_P700 = 0.48
E0_FA = -0.55
E0_Fd = -0.43
E0_NADP = -0.113
convf = 0.032
R = 0.0083
T = 298
Carotenoids_tot = 1
Fd_star = 5
PC_tot = 4
PSBS_tot = 1
LHC_tot = 1
gamma0 = 0.06260060801266355
gamma1 = 0.4053583123566203
gamma2 = 0.7040758738825375
gamma3 = 0.07834807781016208
kZSat = 0.12
E0_QA = -0.14
E0_PQ = 0.354
PQ_tot = 17.5
staticAntII = 0.1
staticAntI = 0.37
Thioredoxin_tot = 1
E_total = 6
NADP_star = 0.8
A_star_P = 2.55
Pi_tot = 17.05
kf_ferredoxin_thioredoxin_reductase = 0.8
kf_tr_activation = 1
kf_tr_inactivation = 0.1
ASC_tot_star = 10
Glutathion_tot = 10
kf_atp_synthase = 20
HPR = 4.666666666666667
Pi_mol = 0.01
DeltaG0_ATP = 30.6
kh_lhc_protonation = 10
kf_lhc_protonation = 0.15837051384170664
ksat_lhc_protonation = 6.2539066418842255
kf_lhc_deprotonation = 0.015892570403695704
kf_cyclic_electron_flow = 1
kf_violaxanthin_deepoxidase = 0.0006091912188339879
kh_violaxanthin_deepoxidase = 4
ksat_violaxanthin_deepoxidase = 6.193595407850397
kf_zeaxanthin_epoxidase = 0.000106261953934132
km_fnr_Ferredoxine_reduced = 1.56
km_fnr_NADP = 0.22
E0_fnr = 3
kcat_fnr = 500
kf_ndh = 0.002
PSII_total = 2.5
PSI_total = 2.5
kH0 = 500000000
kPQred = 250
kPCox = 2500
kFdred = 250000
k2 = 5000000000
kH = 5000000000
kF = 625000000
kMehler = 1
kf_proton_leak = 10
kPTOX = 0.01
kStt7 = 0.0035
km_lhc_state_transition_12 = 0.2
n_ST = 2
kPph1 = 0.0013
E0_rubisco = 1
kcat_rubisco_carboxylase = 2.72
km_rubisco_carboxylase_RUBP = 0.02
km_rubisco_carboxylase_CO2_dissolved = 0.0107
ki_rubisco_carboxylase_3PGA = 0.04
ki_rubisco_carboxylase_FBP = 0.04
ki_rubisco_carboxylase_SBP = 0.075
ki_rubisco_carboxylase_Orthophosphate = 0.9
ki_rubisco_carboxylase_NADPH = 0.07
kre_phosphoglycerate_kinase = 800000000
keq_phosphoglycerate_kinase = 0.00031
kre_gadph = 800000000
keq_gadph = 16000000
kre_triose_phosphate_isomerase = 800000000
keq_triose_phosphate_isomerase = 22
kre_aldolase_dhap_gap = 800000000
keq_aldolase_dhap_gap = 7.1
kre_aldolase_dhap_e4p = 800000000
keq_aldolase_dhap_e4p = 13
E0_fbpase = 1
kcat_fbpase = 1.6
km_fbpase_s = 0.03
ki_fbpase_F6P = 0.7
ki_fbpase_Orthophosphate = 12
kre_transketolase_gap_f6p = 800000000
keq_transketolase_gap_f6p = 0.084
kre_transketolase_gap_s7p = 800000000
keq_transketolase_gap_s7p = 0.85
E0_SBPase = 1
kcat_SBPase = 0.32
km_SBPase_s = 0.013
ki_SBPase_Orthophosphate = 12
kre_ribose_phosphate_isomerase = 800000000
keq_ribose_phosphate_isomerase = 0.4
kre_ribulose_phosphate_epimerase = 800000000
keq_ribulose_phosphate_epimerase = 0.67
E0_phosphoribulokinase = 1
kcat_phosphoribulokinase = 7.9992
km_phosphoribulokinase_RU5P = 0.05
km_phosphoribulokinase_ATP = 0.05
ki_phosphoribulokinase_3PGA = 2
ki_phosphoribulokinase_RUBP = 0.7
ki_phosphoribulokinase_Orthophosphate = 4
ki_phosphoribulokinase_4 = 2.5
ki_phosphoribulokinase_5 = 0.4
kre_g6pi = 800000000
keq_g6pi = 2.3
kre_phosphoglucomutase = 800000000
keq_phosphoglucomutase = 0.058
Orthophosphate_external = 0.5
km_ex_pga = 0.25
km_ex_gap = 0.075
km_ex_dhap = 0.077
km_N_translocator_Orthophosphate_external = 0.74
km_N_translocator_Orthophosphate = 0.63
kcat_N_translocator = 2
E0_N_translocator = 1
E0_ex_g1p = 1
km_ex_g1p_G1P = 0.08
km_ex_g1p_ATP = 0.08
ki_ex_g1p = 10
ki_ex_g1p_3PGA = 0.1
ki_ex_g1p_F6P = 0.02
ki_ex_g1p_FBP = 0.02
kcat_ex_g1p = 0.32
kf_mda_reductase_1 = 500
E0_mda_reductase_2 = 0.002
kcat_mda_reductase_2 = 300
km_mda_reductase_2_NADPH = 0.023
km_mda_reductase_2_MDA = 0.0014
kf1 = 10000
kr1 = 220
kf2 = 10000
kr2 = 4000
kf3 = 2510
kf4 = 10000
kr4 = 4000
kf5 = 2510
XT = 0.07
E0_glutathion_reductase = 0.0014
kcat_glutathion_reductase = 595
km_glutathion_reductase_NADPH = 0.003
km_glutathion_reductase_GSSG = 0.2
km_dehydroascorbate_reductase_DHA = 0.07
km_dehydroascorbate_reductase_GSH = 2.5
K = 0.5
E0_dehydroascorbate_reductase = 0.0017
kcat_dehydroascorbate_reductase = 142
kf_ex_atp = 0.5
kf_ex_nadph = 0.5
kH_Qslope = 30000000000
b6f_content = 1
max_b6f = 500
pKreg = 7
stroma_buffering = 400
kActATPase = 0.001
kDeactATPase = 0.002
k_ATPsynthase = 20
b = 1.8688304401249531
pK0E = 5.960025833706074
k_import_ATP = 0.5
k_import_NADPH = 0.5
volts_per_charge = 0.000769481926574965
ClCe_PQ = 15.87880046767565
Cl_leak_PQ = 14.92901445507139
KEA3_ATP_treshold = 0.26274793681796166
KEA3_pH_reg = 7.69
K_delta_psi_treshold = 0.08146807307624158
VCCN_delta_psi_treshold = 0.08000900979332677
k_Cl_leak = 25
k_NDH1 = 7.447430768265866
k_KEA = 90
perm_K = 1.6113135416150155
k_VCCN1 = 0.5
k_ClCe = 0.5
K_total = 60
Cl_total = 50
ClCe_ATP_threshold = 0.2
RT = R * T
dG_pH = 2.302585092994046 * R * T
Zeaxanthin = Carotenoids_tot + - Violaxanthin
Ferredoxine_reduced = Fd_star + - Ferredoxine_oxidised
Plastocyanine_reduced = PC_tot + - Plastocyanine_oxidised
PsbS_protonated = PSBS_tot + - PsbS_de_minus_protonated
Light_minus_harvesting_complex_protonated = LHC_tot + - Light_minus_harvesting_complex
Q = PsbS_de_minus_protonated * Violaxanthin * gamma0 + PsbS_protonated * Violaxanthin * gamma1 + (PsbS_de_minus_protonated * Zeaxanthin * gamma3) / (Zeaxanthin + kZSat) + (PsbS_protonated * Zeaxanthin * gamma2) / (Zeaxanthin + kZSat)
keq_Plastoquinone_reduced = np.exp((2 * E0_PQ * F + - 2 * E0_QA * F + - 2 * dG_pH * pH) / (RT))
Plastoquinone_reduced = PQ_tot + - Plastoquinone_oxidised
PSII_cross_section = staticAntII + Light_minus_harvesting_complex * (1 + - staticAntI + - staticAntII)
Thioredoxin_reduced = Thioredoxin_tot + - Thioredoxin_oxidised
E_active = E_total + - E_inactive
NADP = NADP_star + - NADPH
ADP = A_star_P + - ATP
Orthophosphate = Pi_tot + - ATP + - DHAP + - E4P + - F6P + - G1P + - G6P + - GAP + - R5P + - RU5P + - S7P + - X5P + - _3PGA + - 2 * BPGA + - 2 * FBP + - 2 * RUBP + - 2 * SBP
ascorbate = ASC_tot_star + - DHA + - MDA
GSH = Glutathion_tot + - 2 * GSSG
keq_atp_synthase = Pi_mol * np.exp((- DeltaG0_ATP + HPR * dG_pH * (pH + - pH_lumen)) / (RT))
keq_fnr = np.exp((- dG_pH * pH + 2 * E0_NADP * F + - 2 * E0_Fd * F) / (RT))
vmax_fnr = E0_fnr * kcat_fnr
E0_rubisco_active = E0_rubisco * E_active
vmax_rubisco_carboxylase = E0_rubisco_active * kcat_rubisco_carboxylase
E0_fbpase_active = E0_fbpase * E_active
vmax_fbpase = E0_fbpase_active * kcat_fbpase
E0_SBPase_active = E0_SBPase * E_active
vmax_SBPase = E0_SBPase_active * kcat_SBPase
E0_phosphoribulokinase_active = E0_phosphoribulokinase * E_active
vmax_phosphoribulokinase = E0_phosphoribulokinase_active * kcat_phosphoribulokinase
vmax_ex_pga = E0_N_translocator * kcat_N_translocator
N_translocator = 1 + (1 + (km_N_translocator_Orthophosphate_external) / (Orthophosphate_external)) * ((DHAP) / (km_ex_dhap) + (GAP) / (km_ex_gap) + (Orthophosphate) / (km_N_translocator_Orthophosphate) + (_3PGA) / (km_ex_pga))
E0_ex_g1p_active = E0_ex_g1p * E_active
vmax_ex_g1p = E0_ex_g1p_active * kcat_ex_g1p
vmax_mda_reductase_2 = E0_mda_reductase_2 * kcat_mda_reductase_2
vmax_glutathion_reductase = E0_glutathion_reductase * kcat_glutathion_reductase
vmax_dehydroascorbate_reductase = E0_dehydroascorbate_reductase * kcat_dehydroascorbate_reductase
keq_PCP700 = np.exp((E0_P700 * F + - E0_PC * F) / (RT))
keq_FAFd = np.exp((E0_Fd * F + - E0_FA * F) / (RT))
B3 = PSII_total + - B0 + - B1 + - B2
P700_plus_FA = PSI_total + - P700FA + - P700FA_minus + - P700_plus_FA_minus
rel_P700_plus_FA = (P700_plus_FA) / (PSI_total)
rel_P700FA = (P700FA) / (PSI_total)
rel_P700FA_minus = (P700FA_minus) / (PSI_total)
rel_P700_plus_FA_minus = (P700_plus_FA_minus) / (PSI_total)
rel_P700 = (P700_plus_FA + P700_plus_FA_minus) / (PSI_total)
rel_P700_plus = (P700_plus_FA + P700_plus_FA_minus) / (PSI_total)
rel_B0 = (B0) / (PSII_total)
rel_B1 = (B1) / (PSII_total)
rel_B2 = (B2) / (PSII_total)
rel_B3 = (B3) / (PSII_total)
Fluo = (B0 * PSII_cross_section * kF) / (k2 + kF + kH0 + Q * kH_Qslope) + (B2 * PSII_cross_section * kF) / (kF + kH0 + Q * kH_Qslope)
keq_b6f_dyn = b6f_content * max_b6f * (1 + - (1) / (1 + (10) ** (pH_lumen + - pKreg)))
protons_lumen = 4000 * (10) ** (- pH_lumen)
protons = 31250 * (10) ** (- pH)
ATP_pmf_activity = ((10) ** (- pK0E) * np.exp((F * b * (delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F))) / (RT))) / (1 + (10) ** (- pK0E) * np.exp((F * b * (delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F))) / (RT)))
deltapH = dG_pH * (pH + - pH_lumen)
deltapH_in_volts = (2.302585092994046 * R * T * (pH + - pH_lumen)) / (F)
pmf = deltapH + F * delta_psi
pmf_in_V = delta_psi + - (2.302585092994046 * RT * (pH_lumen + - pH)) / (F)
keq_b6f = np.exp((- 2 * pmf_in_V + 2 * E0_PC * F + 2 * dG_pH * pH_lumen + - 2 * E0_PQ * F) / (RT))
K_lumen = K_total + - K_stroma
Cl_lumen = Cl_total + - Cl_stroma
total_Cl_2 = (Cl_total) ** (2)
total_K_2 = (K_total) ** (2)
KEA3_reg = (0.81) / ((1 + np.exp(100 * KEA3_ATP_treshold + - 100 * ATP)) * (1 + np.exp(1000 * pH + - 1000 * KEA3_pH_reg)))
dG_K_ions = F * (delta_psi + - (RT * np.log((K_stroma) / (K_lumen))) / (F * np.log(10)))
Cl_driving_force = F * (delta_psi + (RT * np.log((Cl_stroma) / (Cl_lumen))) / (F * np.log(10)))
Keq_NDH1 = np.exp((- 4 * pmf + 2 * E0_PQ * F + - 2 * E0_Fd * F + - 2 * dG_pH * pH) / (RT))
ClCe_activation = (0.9) / (1 + np.exp(100 * ATP + - 100 * ClCe_ATP_threshold))
PQ_ox_div_tot = (Plastoquinone_reduced) / (PQ_tot)
Fd_ox_div_tot = (Ferredoxine_reduced) / (Fd_star)
PC_ox_div_tot = (Plastocyanine_reduced) / (PC_tot)
NADPH_div_tot = (NADPH) / (NADP_star)
ATP_div_tot = (ATP) / (A_star_P)
ferredoxin_thioredoxin_reductase = Ferredoxine_reduced * Thioredoxin_oxidised * kf_ferredoxin_thioredoxin_reductase
tr_activation = E_inactive * Thioredoxin_reduced * kf_tr_activation
tr_inactivation = E_active * kf_tr_inactivation
atp_synthase = ATP_pmf_activity * ATPactivity * k_ATPsynthase * ((ADP) / (convf) + - (ATP) / (convf * kf_atp_synthase))
lhc_protonation = (PsbS_de_minus_protonated * kf_lhc_protonation * (protons_lumen) ** (kh_lhc_protonation)) / ((protons_lumen) ** (kh_lhc_protonation) + (4000 * (10) ** (- ksat_lhc_protonation)) ** (kh_lhc_protonation))
lhc_deprotonation = PsbS_protonated * kf_lhc_deprotonation
cyclic_electron_flow = Plastoquinone_oxidised * kf_cyclic_electron_flow * (Ferredoxine_reduced) ** (2)
violaxanthin_deepoxidase = (Violaxanthin * kf_violaxanthin_deepoxidase * (protons_lumen) ** (kh_violaxanthin_deepoxidase)) / ((protons_lumen) ** (kh_violaxanthin_deepoxidase) + (4000 * (10) ** (- ksat_violaxanthin_deepoxidase)) ** (kh_violaxanthin_deepoxidase))
zeaxanthin_epoxidase = Zeaxanthin * kf_zeaxanthin_epoxidase
fnr = (vmax_fnr * ((NADP * ((Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) ** (2)) / (convf * km_fnr_NADP) + - (NADPH * ((Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)) ** (2)) / (convf * keq_fnr * km_fnr_NADP))) / (-1 + (1 + (NADP) / (convf * km_fnr_NADP)) * (1 + ((Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) ** (2) + (Ferredoxine_reduced) / (km_fnr_Ferredoxine_reduced)) + (1 + (NADPH) / (convf * km_fnr_NADP)) * (1 + ((Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)) ** (2) + (Ferredoxine_oxidised) / (km_fnr_Ferredoxine_reduced)))
ndh = Plastoquinone_oxidised * kf_ndh
proton_leak = kf_proton_leak * (protons_lumen + - 4000 * (10) ** (- pH))
PTOX = O2_dissolved_lumen * Plastoquinone_reduced * kPTOX
lhc_state_transition_12 = (1 * Light_minus_harvesting_complex * kStt7) / (1 + ((Plastoquinone_oxidised) / (PQ_tot * km_lhc_state_transition_12)) ** (n_ST))
lhc_state_transition_21 = Light_minus_harvesting_complex_protonated * kPph1
rubisco_carboxylase = (CO2_dissolved * RUBP * vmax_rubisco_carboxylase) / ((CO2_dissolved + km_rubisco_carboxylase_CO2_dissolved) * (RUBP + km_rubisco_carboxylase_RUBP * (1 + (FBP) / (ki_rubisco_carboxylase_FBP) + (NADPH) / (ki_rubisco_carboxylase_NADPH) + (Orthophosphate) / (ki_rubisco_carboxylase_Orthophosphate) + (SBP) / (ki_rubisco_carboxylase_SBP) + (_3PGA) / (ki_rubisco_carboxylase_3PGA))))
phosphoglycerate_kinase = kre_phosphoglycerate_kinase * (ATP * _3PGA + - (ADP * BPGA) / (keq_phosphoglycerate_kinase))
gadph = kre_gadph * (BPGA * NADPH * protons + - (GAP * NADP * Orthophosphate) / (keq_gadph))
triose_phosphate_isomerase = kre_triose_phosphate_isomerase * (GAP + - (DHAP) / (keq_triose_phosphate_isomerase))
aldolase_dhap_gap = kre_aldolase_dhap_gap * (DHAP * GAP + - (FBP) / (keq_aldolase_dhap_gap))
aldolase_dhap_e4p = kre_aldolase_dhap_e4p * (DHAP * E4P + - (SBP) / (keq_aldolase_dhap_e4p))
fbpase = (FBP * vmax_fbpase) / (FBP + km_fbpase_s * (1 + (F6P) / (ki_fbpase_F6P) + (Orthophosphate) / (ki_fbpase_Orthophosphate)))
transketolase_gap_f6p = kre_transketolase_gap_f6p * (F6P * GAP + - (E4P * X5P) / (keq_transketolase_gap_f6p))
transketolase_gap_s7p = kre_transketolase_gap_s7p * (GAP * S7P + - (R5P * X5P) / (keq_transketolase_gap_s7p))
SBPase = (SBP * vmax_SBPase) / (SBP + km_SBPase_s * (1 + (Orthophosphate) / (ki_SBPase_Orthophosphate)))
ribose_phosphate_isomerase = kre_ribose_phosphate_isomerase * (R5P + - (RU5P) / (keq_ribose_phosphate_isomerase))
ribulose_phosphate_epimerase = kre_ribulose_phosphate_epimerase * (X5P + - (RU5P) / (keq_ribulose_phosphate_epimerase))
phosphoribulokinase = (ATP * RU5P * vmax_phosphoribulokinase) / ((RU5P + km_phosphoribulokinase_RU5P * (1 + (Orthophosphate) / (ki_phosphoribulokinase_Orthophosphate) + (RUBP) / (ki_phosphoribulokinase_RUBP) + (_3PGA) / (ki_phosphoribulokinase_3PGA))) * (ATP * (1 + (ADP) / (ki_phosphoribulokinase_4)) + km_phosphoribulokinase_ATP * (1 + (ADP) / (ki_phosphoribulokinase_5))))
g6pi = kre_g6pi * (F6P + - (G6P) / (keq_g6pi))
phosphoglucomutase = kre_phosphoglucomutase * (G6P + - (G1P) / (keq_phosphoglucomutase))
ex_pga = (_3PGA * vmax_ex_pga) / (N_translocator * km_ex_pga)
ex_gap = (GAP * vmax_ex_pga) / (N_translocator * km_ex_gap)
ex_dhap = (DHAP * vmax_ex_pga) / (N_translocator * km_ex_dhap)
ex_g1p = (ATP * G1P * vmax_ex_g1p) / ((G1P + km_ex_g1p_G1P) * ((1 + (ADP) / (ki_ex_g1p)) * (ATP + km_ex_g1p_ATP) + (Orthophosphate * km_ex_g1p_ATP) / (F6P * ki_ex_g1p_F6P + FBP * ki_ex_g1p_FBP + _3PGA * ki_ex_g1p_3PGA)))
mda_reductase_1 = kf_mda_reductase_1 * (MDA) ** (2)
mda_reductase_2 = (MDA * NADPH * vmax_mda_reductase_2) / (MDA * NADPH + MDA * km_mda_reductase_2_NADPH + NADPH * km_mda_reductase_2_MDA + km_mda_reductase_2_MDA * km_mda_reductase_2_NADPH)
ascorbate_peroxidase = (H2O2 * XT * ascorbate) / ((H2O2) / (kf2) + (H2O2) / (kf4) + (ascorbate) / (kf1) + H2O2 * ascorbate * ((1) / (kf3) + (1) / (kf5)) + (kr1) / (kf1 * kf2) + (H2O2 * kr2) / (kf2 * kf3) + (H2O2 * kr4) / (kf4 * kf5) + (kr1 * kr2) / (kf1 * kf2 * kf3))
glutathion_reductase = (GSSG * NADPH * vmax_glutathion_reductase) / (GSSG * NADPH + GSSG * km_glutathion_reductase_NADPH + NADPH * km_glutathion_reductase_GSSG + km_glutathion_reductase_GSSG * km_glutathion_reductase_NADPH)
dehydroascorbate_reductase = (DHA * GSH * vmax_dehydroascorbate_reductase) / (K + DHA * GSH + DHA * km_dehydroascorbate_reductase_GSH + GSH * km_dehydroascorbate_reductase_DHA)
toP700FA_minus = P700_plus_FA_minus * Plastocyanine_reduced * kPCox + - (P700FA_minus * Plastocyanine_oxidised * kPCox) / (keq_PCP700)
toP700FA_v3 = Ferredoxine_oxidised * P700FA_minus * kFdred + - (Ferredoxine_reduced * P700FA * kFdred) / (keq_FAFd)
toP700_plus_FA = Ferredoxine_oxidised * P700_plus_FA_minus * kFdred + - (Ferredoxine_reduced * P700_plus_FA * kFdred) / (keq_FAFd)
toP700FA_v5 = P700_plus_FA * Plastocyanine_reduced * kPCox + - (P700FA * Plastocyanine_oxidised * kPCox) / (keq_PCP700)
PSI = P700FA * PPFD * (1 + - PSII_cross_section)
mehler1 = O2_dissolved_lumen * P700FA_minus * kMehler
mehler2 = O2_dissolved_lumen * P700_plus_FA_minus * kMehler
B01 = B0 * PPFD * PSII_cross_section
B10Q = B1 * (kH0 + Q * kH_Qslope)
B10F = B1 * kF
B12 = B1 * k2
B20 = B2 * Plastoquinone_oxidised * kPQred + - (B0 * Plastoquinone_reduced * kPQred) / (keq_Plastoquinone_reduced)
B23 = B2 * PPFD * PSII_cross_section
B32F = B3 * kF
B32Q = B3 * (kH0 + Q * kH_Qslope)
b6f = (Plastocyanine_oxidised * Plastoquinone_reduced * keq_b6f_dyn) / (Plastoquinone_oxidised + Plastoquinone_reduced) + - (Plastocyanine_reduced * keq_b6f_dyn * (1 + - (Plastoquinone_reduced) / (Plastoquinone_oxidised + Plastoquinone_reduced))) / (keq_b6f)
vATPactivity = kActATPase * (1 + - ATPactivity) if PPFD > 0 else - ATPactivity * kDeactATPase
vATP_shuttle = ADP * k_import_ATP + - ATP * kf_ex_atp
vNADPH_shuttle = NADP * k_import_NADPH + - NADPH * kf_ex_nadph
KEA3 = max(0, KEA3_reg * k_KEA * (K_stroma * protons_lumen + - K_lumen * protons))
voltage_K_channel = (0.9 * K_lumen * dG_K_ions * perm_K) / (K_stroma * (1 + np.exp(1000 * K_delta_psi_treshold + - 1000 * delta_psi)))
VCCN1 = (0.9 * Cl_driving_force * Cl_stroma * k_VCCN1) / (Cl_lumen * (1 + np.exp(1000 * VCCN_delta_psi_treshold + - 1000 * delta_psi)))
Cl_leak = (0.9 * k_Cl_leak * (Cl_lumen + - Cl_stroma) ** (2)) / (total_Cl_2 * (1 + np.exp(10 * Cl_leak_PQ + - 10 * Plastoquinone_oxidised)))
NDH1 = (0.9 * Plastoquinone_oxidised * k_NDH1 * (10) ** (-6.5 + pH_lumen) * (Ferredoxine_reduced) ** (2)) / ((0.5 + (10) ** (-6.5 + pH_lumen)) * (1 + 7.38905609893065 * np.exp(- 100 * P700_plus_FA_minus)))
ClCe_bi = ClCe_activation * k_ClCe * (Cl_stroma + - Cl_lumen)
return [RT, dG_pH, Zeaxanthin, Ferredoxine_reduced, Plastocyanine_reduced, PsbS_protonated, Light_minus_harvesting_complex_protonated, Q, keq_Plastoquinone_reduced, Plastoquinone_reduced, PSII_cross_section, Thioredoxin_reduced, E_active, NADP, ADP, Orthophosphate, ascorbate, GSH, keq_atp_synthase, keq_fnr, vmax_fnr, E0_rubisco_active, vmax_rubisco_carboxylase, E0_fbpase_active, vmax_fbpase, E0_SBPase_active, vmax_SBPase, E0_phosphoribulokinase_active, vmax_phosphoribulokinase, vmax_ex_pga, N_translocator, E0_ex_g1p_active, vmax_ex_g1p, vmax_mda_reductase_2, vmax_glutathion_reductase, vmax_dehydroascorbate_reductase, keq_PCP700, keq_FAFd, B3, P700_plus_FA, rel_P700_plus_FA, rel_P700FA, rel_P700FA_minus, rel_P700_plus_FA_minus, rel_P700, rel_P700_plus, rel_B0, rel_B1, rel_B2, rel_B3, Fluo, keq_b6f_dyn, protons_lumen, protons, ATP_pmf_activity, deltapH, deltapH_in_volts, pmf, pmf_in_V, keq_b6f, K_lumen, Cl_lumen, total_Cl_2, total_K_2, KEA3_reg, dG_K_ions, Cl_driving_force, Keq_NDH1, ClCe_activation, PQ_ox_div_tot, Fd_ox_div_tot, PC_ox_div_tot, NADPH_div_tot, ATP_div_tot, ferredoxin_thioredoxin_reductase, tr_activation, tr_inactivation, atp_synthase, lhc_protonation, lhc_deprotonation, cyclic_electron_flow, violaxanthin_deepoxidase, zeaxanthin_epoxidase, fnr, ndh, proton_leak, PTOX, lhc_state_transition_12, lhc_state_transition_21, rubisco_carboxylase, phosphoglycerate_kinase, gadph, triose_phosphate_isomerase, aldolase_dhap_gap, aldolase_dhap_e4p, fbpase, transketolase_gap_f6p, transketolase_gap_s7p, SBPase, ribose_phosphate_isomerase, ribulose_phosphate_epimerase, phosphoribulokinase, g6pi, phosphoglucomutase, ex_pga, ex_gap, ex_dhap, ex_g1p, mda_reductase_1, mda_reductase_2, ascorbate_peroxidase, glutathion_reductase, dehydroascorbate_reductase, toP700FA_minus, toP700FA_v3, toP700_plus_FA, toP700FA_v5, PSI, mehler1, mehler2, B01, B10Q, B10F, B12, B20, B23, B32F, B32Q, b6f, vATPactivity, vATP_shuttle, vNADPH_shuttle, KEA3, voltage_K_channel, VCCN1, Cl_leak, NDH1, ClCe_bi]
derived = all_derived
y0 = {"_3PGA": 0.9167729479368978, "BPGA": 0.0003814495319659031, "GAP": 0.00580821050261484, "DHAP": 0.1277806166216142, "FBP": 0.005269452472931973, "F6P": 0.2874944558066638, "G6P": 0.6612372482712676, "G1P": 0.03835176039761378, "SBP": 0.011101373736607443, "S7P": 0.1494578301900007, "E4P": 0.00668295494870102, "X5P": 0.020988553174809618, "R5P": 0.035155825913785584, "RUBP": 0.11293260727162346, "RU5P": 0.014062330254191594, "ATP": 1.4612747767895344, "Ferredoxine_oxidised": 3.715702384326767, "Light_minus_harvesting_complex": 0.7805901436176024, "NADPH": 0.5578718406315588, "Plastocyanine_oxidised": 1.8083642974980014, "Plastoquinone_oxidised": 10.251099271612473, "PsbS_de_minus_protonated": 0.9667381262477079, "Violaxanthin": 0.9629870646993118, "MDA": 2.0353396709300447e-7, "H2O2": 1.2034405327140102e-7, "DHA": 1.0296456279861962e-11, "GSSG": 4.99986167652437e-12, "Thioredoxin_oxidised": 0.9334426859846461, "E_inactive": 3.6023635680406634, "P700FA": 1.506615384275408, "P700_plus_FA_minus": 0.019197449388051676, "P700FA_minus": 0.028144516332212766, "B0": 1.9379789566530539, "B1": 9.786232812526368e-8, "B2": 0.5620208537555176, "pH_lumen": 6.8, "pH": 7.5, "ATPactivity": 0, "delta_psi": 0, "K_stroma": 30, "Cl_stroma": 25}
Generated LaTeX Code
\begin{align*}
\frac{d 3PGA}{dt} &= 2 \cdot \frac{CO2 (dissolved) \cdot RUBP \cdot vmax\_rubisco\_carboxylase}{(CO2 (dissolved) + km\_rubisco\_carboxylase\_CO2 (dissolved)) \cdot (RUBP + km\_rubisco\_carboxylase\_RUBP \cdot (1 + \frac{FBP}{ki\_rubisco\_carboxylase\_FBP} + \frac{NADPH}{ki\_rubisco\_carboxylase\_NADPH} + \frac{Orthophosphate}{ki\_rubisco\_carboxylase\_Orthophosphate} + \frac{SBP}{ki\_rubisco\_carboxylase\_SBP} + \frac{3PGA}{ki\_rubisco\_carboxylase\_3PGA}))} \\
& - kre\_phosphoglycerate\_kinase \cdot (ATP \cdot 3PGA - \frac{ADP \cdot BPGA}{keq\_phosphoglycerate\_kinase}) \\
& - \frac{3PGA \cdot vmax\_ex\_pga}{N\_translocator \cdot km\_ex\_pga}\\
\frac{d BPGA}{dt} &= kre\_phosphoglycerate\_kinase \cdot (ATP \cdot 3PGA - \frac{ADP \cdot BPGA}{keq\_phosphoglycerate\_kinase}) \\
& - kre\_gadph \cdot (BPGA \cdot NADPH \cdot protons - \frac{GAP \cdot NADP \cdot Orthophosphate}{keq\_gadph})\\
\frac{d GAP}{dt} &= kre\_gadph \cdot (BPGA \cdot NADPH \cdot protons - \frac{GAP \cdot NADP \cdot Orthophosphate}{keq\_gadph}) \\
& - kre\_triose\_phosphate\_isomerase \cdot (GAP - \frac{DHAP}{keq\_triose\_phosphate\_isomerase}) \\
& - kre\_aldolase\_dhap\_gap \cdot (DHAP \cdot GAP - \frac{FBP}{keq\_aldolase\_dhap\_gap}) \\
& - kre\_transketolase\_gap\_f6p \cdot (F6P \cdot GAP - \frac{E4P \cdot X5P}{keq\_transketolase\_gap\_f6p}) \\
& - kre\_transketolase\_gap\_s7p \cdot (GAP \cdot S7P - \frac{R5P \cdot X5P}{keq\_transketolase\_gap\_s7p}) \\
& - \frac{GAP \cdot vmax\_ex\_pga}{N\_translocator \cdot km\_ex\_gap}\\
\frac{d DHAP}{dt} &= kre\_triose\_phosphate\_isomerase \cdot (GAP - \frac{DHAP}{keq\_triose\_phosphate\_isomerase}) \\
& - kre\_aldolase\_dhap\_gap \cdot (DHAP \cdot GAP - \frac{FBP}{keq\_aldolase\_dhap\_gap}) \\
& - kre\_aldolase\_dhap\_e4p \cdot (DHAP \cdot E4P - \frac{SBP}{keq\_aldolase\_dhap\_e4p}) \\
& - \frac{DHAP \cdot vmax\_ex\_pga}{N\_translocator \cdot km\_ex\_dhap}\\
\frac{d FBP}{dt} &= kre\_aldolase\_dhap\_gap \cdot (DHAP \cdot GAP - \frac{FBP}{keq\_aldolase\_dhap\_gap}) \\
& - \frac{FBP \cdot vmax\_fbpase}{FBP + km\_fbpase\_s \cdot (1 + \frac{F6P}{ki\_fbpase\_F6P} + \frac{Orthophosphate}{ki\_fbpase\_Orthophosphate})}\\
\frac{d F6P}{dt} &= \frac{FBP \cdot vmax\_fbpase}{FBP + km\_fbpase\_s \cdot (1 + \frac{F6P}{ki\_fbpase\_F6P} + \frac{Orthophosphate}{ki\_fbpase\_Orthophosphate})} \\
& - kre\_transketolase\_gap\_f6p \cdot (F6P \cdot GAP - \frac{E4P \cdot X5P}{keq\_transketolase\_gap\_f6p}) \\
& - kre\_g6pi \cdot (F6P - \frac{G6P}{keq\_g6pi})\\
\frac{d G6P}{dt} &= kre\_g6pi \cdot (F6P - \frac{G6P}{keq\_g6pi}) \\
& - kre\_phosphoglucomutase \cdot (G6P - \frac{G1P}{keq\_phosphoglucomutase})\\
\frac{d G1P}{dt} &= kre\_phosphoglucomutase \cdot (G6P - \frac{G1P}{keq\_phosphoglucomutase}) \\
& - \frac{ATP \cdot G1P \cdot vmax\_ex\_g1p}{(G1P + km\_ex\_g1p\_G1P) \cdot ((1 + \frac{ADP}{ki\_ex\_g1p}) \cdot (ATP + km\_ex\_g1p\_ATP) + \frac{Orthophosphate \cdot km\_ex\_g1p\_ATP}{F6P \cdot ki\_ex\_g1p\_F6P + FBP \cdot ki\_ex\_g1p\_FBP + 3PGA \cdot ki\_ex\_g1p\_3PGA})}\\
\frac{d SBP}{dt} &= kre\_aldolase\_dhap\_e4p \cdot (DHAP \cdot E4P - \frac{SBP}{keq\_aldolase\_dhap\_e4p}) \\
& - \frac{SBP \cdot vmax\_SBPase}{SBP + km\_SBPase\_s \cdot (1 + \frac{Orthophosphate}{ki\_SBPase\_Orthophosphate})}\\
\frac{d S7P}{dt} &= - kre\_transketolase\_gap\_s7p \cdot (GAP \cdot S7P - \frac{R5P \cdot X5P}{keq\_transketolase\_gap\_s7p}) \\
& + \frac{SBP \cdot vmax\_SBPase}{SBP + km\_SBPase\_s \cdot (1 + \frac{Orthophosphate}{ki\_SBPase\_Orthophosphate})}\\
\frac{d E4P}{dt} &= - kre\_aldolase\_dhap\_e4p \cdot (DHAP \cdot E4P - \frac{SBP}{keq\_aldolase\_dhap\_e4p}) \\
& + kre\_transketolase\_gap\_f6p \cdot (F6P \cdot GAP - \frac{E4P \cdot X5P}{keq\_transketolase\_gap\_f6p})\\
\frac{d X5P}{dt} &= kre\_transketolase\_gap\_f6p \cdot (F6P \cdot GAP - \frac{E4P \cdot X5P}{keq\_transketolase\_gap\_f6p}) \\
& + kre\_transketolase\_gap\_s7p \cdot (GAP \cdot S7P - \frac{R5P \cdot X5P}{keq\_transketolase\_gap\_s7p}) \\
& - kre\_ribulose\_phosphate\_epimerase \cdot (X5P - \frac{RU5P}{keq\_ribulose\_phosphate\_epimerase})\\
\frac{d R5P}{dt} &= kre\_transketolase\_gap\_s7p \cdot (GAP \cdot S7P - \frac{R5P \cdot X5P}{keq\_transketolase\_gap\_s7p}) \\
& - kre\_ribose\_phosphate\_isomerase \cdot (R5P - \frac{RU5P}{keq\_ribose\_phosphate\_isomerase})\\
\frac{d RUBP}{dt} &= - \frac{CO2 (dissolved) \cdot RUBP \cdot vmax\_rubisco\_carboxylase}{(CO2 (dissolved) + km\_rubisco\_carboxylase\_CO2 (dissolved)) \cdot (RUBP + km\_rubisco\_carboxylase\_RUBP \cdot (1 + \frac{FBP}{ki\_rubisco\_carboxylase\_FBP} + \frac{NADPH}{ki\_rubisco\_carboxylase\_NADPH} + \frac{Orthophosphate}{ki\_rubisco\_carboxylase\_Orthophosphate} + \frac{SBP}{ki\_rubisco\_carboxylase\_SBP} + \frac{3PGA}{ki\_rubisco\_carboxylase\_3PGA}))} \\
& + \frac{ATP \cdot RU5P \cdot vmax\_phosphoribulokinase}{(RU5P + km\_phosphoribulokinase\_RU5P \cdot (1 + \frac{Orthophosphate}{ki\_phosphoribulokinase\_Orthophosphate} + \frac{RUBP}{ki\_phosphoribulokinase\_RUBP} + \frac{3PGA}{ki\_phosphoribulokinase\_3PGA})) \cdot (ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_4}) + km\_phosphoribulokinase\_ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_5}))}\\
\frac{d RU5P}{dt} &= kre\_ribose\_phosphate\_isomerase \cdot (R5P - \frac{RU5P}{keq\_ribose\_phosphate\_isomerase}) \\
& + kre\_ribulose\_phosphate\_epimerase \cdot (X5P - \frac{RU5P}{keq\_ribulose\_phosphate\_epimerase}) \\
& - \frac{ATP \cdot RU5P \cdot vmax\_phosphoribulokinase}{(RU5P + km\_phosphoribulokinase\_RU5P \cdot (1 + \frac{Orthophosphate}{ki\_phosphoribulokinase\_Orthophosphate} + \frac{RUBP}{ki\_phosphoribulokinase\_RUBP} + \frac{3PGA}{ki\_phosphoribulokinase\_3PGA})) \cdot (ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_4}) + km\_phosphoribulokinase\_ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_5}))}\\
\frac{d ATP}{dt} &= convf \cdot ATP\_pmf\_activity \cdot ATPactivity \cdot k\_ATPsynthase \cdot (\frac{ADP}{convf} - \frac{ATP}{convf \cdot kf\_atp\_synthase}) \\
& - kre\_phosphoglycerate\_kinase \cdot (ATP \cdot 3PGA - \frac{ADP \cdot BPGA}{keq\_phosphoglycerate\_kinase}) \\
& - \frac{ATP \cdot RU5P \cdot vmax\_phosphoribulokinase}{(RU5P + km\_phosphoribulokinase\_RU5P \cdot (1 + \frac{Orthophosphate}{ki\_phosphoribulokinase\_Orthophosphate} + \frac{RUBP}{ki\_phosphoribulokinase\_RUBP} + \frac{3PGA}{ki\_phosphoribulokinase\_3PGA})) \cdot (ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_4}) + km\_phosphoribulokinase\_ATP \cdot (1 + \frac{ADP}{ki\_phosphoribulokinase\_5}))} \\
& - \frac{ATP \cdot G1P \cdot vmax\_ex\_g1p}{(G1P + km\_ex\_g1p\_G1P) \cdot ((1 + \frac{ADP}{ki\_ex\_g1p}) \cdot (ATP + km\_ex\_g1p\_ATP) + \frac{Orthophosphate \cdot km\_ex\_g1p\_ATP}{F6P \cdot ki\_ex\_g1p\_F6P + FBP \cdot ki\_ex\_g1p\_FBP + 3PGA \cdot ki\_ex\_g1p\_3PGA})} \\
& + ADP \cdot k\_import\_ATP - ATP \cdot kf\_ex\_atp\\
\frac{d Ferredoxine (oxidised)}{dt} &= Ferredoxine (reduced) \cdot Thioredoxin (oxidised) \cdot kf\_ferredoxin\_thioredoxin\_reductase \\
& + 2 \cdot Plastoquinone (oxidised) \cdot kf\_cyclic\_electron\_flow \cdot {Ferredoxine (reduced)}^{2} \\
& + 2 \cdot \frac{vmax\_fnr \cdot (\frac{NADP \cdot {\frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}}^{2}}{convf \cdot km\_fnr\_NADP} - \frac{NADPH \cdot {\frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)}}^{2}}{convf \cdot keq\_fnr \cdot km\_fnr\_NADP})}{-1 + (1 + \frac{NADP}{convf \cdot km\_fnr\_NADP}) \cdot (1 + {\frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}}^{2} + \frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}) + (1 + \frac{NADPH}{convf \cdot km\_fnr\_NADP}) \cdot (1 + {\frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)}}^{2} + \frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)})} \\
& - Ferredoxine (oxidised) \cdot P700FA- \cdot kFdred - \frac{Ferredoxine (reduced) \cdot P700FA \cdot kFdred}{keq\_FAFd} \\
& - Ferredoxine (oxidised) \cdot P700+FA- \cdot kFdred - \frac{Ferredoxine (reduced) \cdot P700+FA \cdot kFdred}{keq\_FAFd} \\
& + 2 \cdot \frac{0.9 \cdot Plastoquinone (oxidised) \cdot k\_NDH1 \cdot {10}^{-6.5 + pH\_lumen} \cdot {Ferredoxine (reduced)}^{2}}{(0.5 + {10}^{-6.5 + pH\_lumen}) \cdot (1 + 7.38905609893065 \cdot e^{- 100 \cdot P700+FA-})}\\
\frac{d Light-harvesting complex}{dt} &= - \frac{1 \cdot Light-harvesting complex \cdot kStt7}{1 + {\frac{Plastoquinone (oxidised)}{PQ\_tot \cdot km\_lhc\_state\_transition\_12}}^{n\_ST}} \\
& + Light-harvesting complex (protonated) \cdot kPph1\\
\frac{d NADPH}{dt} &= convf \cdot \frac{vmax\_fnr \cdot (\frac{NADP \cdot {\frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}}^{2}}{convf \cdot km\_fnr\_NADP} - \frac{NADPH \cdot {\frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)}}^{2}}{convf \cdot keq\_fnr \cdot km\_fnr\_NADP})}{-1 + (1 + \frac{NADP}{convf \cdot km\_fnr\_NADP}) \cdot (1 + {\frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}}^{2} + \frac{Ferredoxine (reduced)}{km\_fnr\_Ferredoxine (reduced)}) + (1 + \frac{NADPH}{convf \cdot km\_fnr\_NADP}) \cdot (1 + {\frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)}}^{2} + \frac{Ferredoxine (oxidised)}{km\_fnr\_Ferredoxine (reduced)})} \\
& - kre\_gadph \cdot (BPGA \cdot NADPH \cdot protons - \frac{GAP \cdot NADP \cdot Orthophosphate}{keq\_gadph}) \\
& - \frac{MDA \cdot NADPH \cdot vmax\_mda\_reductase\_2}{MDA \cdot NADPH + MDA \cdot km\_mda\_reductase\_2\_NADPH + NADPH \cdot km\_mda\_reductase\_2\_MDA + km\_mda\_reductase\_2\_MDA \cdot km\_mda\_reductase\_2\_NADPH} \\
& - \frac{GSSG \cdot NADPH \cdot vmax\_glutathion\_reductase}{GSSG \cdot NADPH + GSSG \cdot km\_glutathion\_reductase\_NADPH + NADPH \cdot km\_glutathion\_reductase\_GSSG + km\_glutathion\_reductase\_GSSG \cdot km\_glutathion\_reductase\_NADPH} \\
& + NADP \cdot k\_import\_NADPH - NADPH \cdot kf\_ex\_nadph\\
\frac{d Plastocyanine (oxidised)}{dt} &= P700+FA- \cdot Plastocyanine (reduced) \cdot kPCox - \frac{P700FA- \cdot Plastocyanine (oxidised) \cdot kPCox}{keq\_PCP700} \\
& + P700+FA \cdot Plastocyanine (reduced) \cdot kPCox - \frac{P700FA \cdot Plastocyanine (oxidised) \cdot kPCox}{keq\_PCP700} \\
& - 2 \cdot \frac{Plastocyanine (oxidised) \cdot Plastoquinone (reduced) \cdot keq\_b6f\_dyn}{Plastoquinone (oxidised) + Plastoquinone (reduced)} - \frac{Plastocyanine (reduced) \cdot keq\_b6f\_dyn \cdot (1 - \frac{Plastoquinone (reduced)}{Plastoquinone (oxidised) + Plastoquinone (reduced)})}{keq\_b6f}\\
\frac{d Plastoquinone (oxidised)}{dt} &= - Plastoquinone (oxidised) \cdot kf\_cyclic\_electron\_flow \cdot {Ferredoxine (reduced)}^{2} \\
& - Plastoquinone (oxidised) \cdot kf\_ndh \\
& + O2 (dissolved)\_lumen \cdot Plastoquinone (reduced) \cdot kPTOX \\
& - 0.5 \cdot B2 \cdot Plastoquinone (oxidised) \cdot kPQred - \frac{B0 \cdot Plastoquinone (reduced) \cdot kPQred}{keq\_Plastoquinone (reduced)} \\
& + \frac{Plastocyanine (oxidised) \cdot Plastoquinone (reduced) \cdot keq\_b6f\_dyn}{Plastoquinone (oxidised) + Plastoquinone (reduced)} - \frac{Plastocyanine (reduced) \cdot keq\_b6f\_dyn \cdot (1 - \frac{Plastoquinone (reduced)}{Plastoquinone (oxidised) + Plastoquinone (reduced)})}{keq\_b6f} \\
& - \frac{0.9 \cdot Plastoquinone (oxidised) \cdot k\_NDH1 \cdot {10}^{-6.5 + pH\_lumen} \cdot {Ferredoxine (reduced)}^{2}}{(0.5 + {10}^{-6.5 + pH\_lumen}) \cdot (1 + 7.38905609893065 \cdot e^{- 100 \cdot P700+FA-})}\\
\frac{d PsbS (de-protonated)}{dt} &= - \frac{PsbS (de-protonated) \cdot kf\_lhc\_protonation \cdot {protons\_lumen}^{kh\_lhc\_protonation}}{{protons\_lumen}^{kh\_lhc\_protonation} + {4000 \cdot {10}^{- ksat\_lhc\_protonation}}^{kh\_lhc\_protonation}} \\
& + PsbS (protonated) \cdot kf\_lhc\_deprotonation\\
\frac{d Violaxanthin}{dt} &= - \frac{Violaxanthin \cdot kf\_violaxanthin\_deepoxidase \cdot {protons\_lumen}^{kh\_violaxanthin\_deepoxidase}}{{protons\_lumen}^{kh\_violaxanthin\_deepoxidase} + {4000 \cdot {10}^{- ksat\_violaxanthin\_deepoxidase}}^{kh\_violaxanthin\_deepoxidase}} \\
& + Zeaxanthin \cdot kf\_zeaxanthin\_epoxidase\\
\frac{d MDA}{dt} &= - 2 \cdot kf\_mda\_reductase\_1 \cdot {MDA}^{2} \\
& - 2 \cdot \frac{MDA \cdot NADPH \cdot vmax\_mda\_reductase\_2}{MDA \cdot NADPH + MDA \cdot km\_mda\_reductase\_2\_NADPH + NADPH \cdot km\_mda\_reductase\_2\_MDA + km\_mda\_reductase\_2\_MDA \cdot km\_mda\_reductase\_2\_NADPH} \\
& + 2 \cdot \frac{H2O2 \cdot XT \cdot ascorbate}{\frac{H2O2}{kf2} + \frac{H2O2}{kf4} + \frac{ascorbate}{kf1} + H2O2 \cdot ascorbate \cdot (\frac{1}{kf3} + \frac{1}{kf5}) + \frac{kr1}{kf1 \cdot kf2} + \frac{H2O2 \cdot kr2}{kf2 \cdot kf3} + \frac{H2O2 \cdot kr4}{kf4 \cdot kf5} + \frac{kr1 \cdot kr2}{kf1 \cdot kf2 \cdot kf3}}\\
\frac{d H2O2}{dt} &= - \frac{H2O2 \cdot XT \cdot ascorbate}{\frac{H2O2}{kf2} + \frac{H2O2}{kf4} + \frac{ascorbate}{kf1} + H2O2 \cdot ascorbate \cdot (\frac{1}{kf3} + \frac{1}{kf5}) + \frac{kr1}{kf1 \cdot kf2} + \frac{H2O2 \cdot kr2}{kf2 \cdot kf3} + \frac{H2O2 \cdot kr4}{kf4 \cdot kf5} + \frac{kr1 \cdot kr2}{kf1 \cdot kf2 \cdot kf3}} \\
& + convf \cdot O2 (dissolved)\_lumen \cdot P700FA- \cdot kMehler \\
& + convf \cdot O2 (dissolved)\_lumen \cdot P700+FA- \cdot kMehler\\
\frac{d DHA}{dt} &= kf\_mda\_reductase\_1 \cdot {MDA}^{2} \\
& - \frac{DHA \cdot GSH \cdot vmax\_dehydroascorbate\_reductase}{K + DHA \cdot GSH + DHA \cdot km\_dehydroascorbate\_reductase\_GSH + GSH \cdot km\_dehydroascorbate\_reductase\_DHA}\\
\frac{d GSSG}{dt} &= - \frac{GSSG \cdot NADPH \cdot vmax\_glutathion\_reductase}{GSSG \cdot NADPH + GSSG \cdot km\_glutathion\_reductase\_NADPH + NADPH \cdot km\_glutathion\_reductase\_GSSG + km\_glutathion\_reductase\_GSSG \cdot km\_glutathion\_reductase\_NADPH} \\
& + \frac{DHA \cdot GSH \cdot vmax\_dehydroascorbate\_reductase}{K + DHA \cdot GSH + DHA \cdot km\_dehydroascorbate\_reductase\_GSH + GSH \cdot km\_dehydroascorbate\_reductase\_DHA}\\
\frac{d Thioredoxin (oxidised)}{dt} &= - Ferredoxine (reduced) \cdot Thioredoxin (oxidised) \cdot kf\_ferredoxin\_thioredoxin\_reductase \\
& + 5 \cdot E\_inactive \cdot Thioredoxin (reduced) \cdot kf\_tr\_activation\\
\frac{d E\_inactive}{dt} &= - 5 \cdot E\_inactive \cdot Thioredoxin (reduced) \cdot kf\_tr\_activation \\
& + 5 \cdot E\_active \cdot kf\_tr\_inactivation\\
\frac{d P700FA}{dt} &= Ferredoxine (oxidised) \cdot P700FA- \cdot kFdred - \frac{Ferredoxine (reduced) \cdot P700FA \cdot kFdred}{keq\_FAFd} \\
& + P700+FA \cdot Plastocyanine (reduced) \cdot kPCox - \frac{P700FA \cdot Plastocyanine (oxidised) \cdot kPCox}{keq\_PCP700} \\
& - P700FA \cdot PPFD \cdot (1 - PSII\_cross\_section) \\
& + 2 \cdot O2 (dissolved)\_lumen \cdot P700FA- \cdot kMehler\\
\frac{d P700+FA-}{dt} &= - P700+FA- \cdot Plastocyanine (reduced) \cdot kPCox - \frac{P700FA- \cdot Plastocyanine (oxidised) \cdot kPCox}{keq\_PCP700} \\
& - Ferredoxine (oxidised) \cdot P700+FA- \cdot kFdred - \frac{Ferredoxine (reduced) \cdot P700+FA \cdot kFdred}{keq\_FAFd} \\
& + P700FA \cdot PPFD \cdot (1 - PSII\_cross\_section) \\
& - 2 \cdot O2 (dissolved)\_lumen \cdot P700+FA- \cdot kMehler\\
\frac{d P700FA-}{dt} &= P700+FA- \cdot Plastocyanine (reduced) \cdot kPCox - \frac{P700FA- \cdot Plastocyanine (oxidised) \cdot kPCox}{keq\_PCP700} \\
& - Ferredoxine (oxidised) \cdot P700FA- \cdot kFdred - \frac{Ferredoxine (reduced) \cdot P700FA \cdot kFdred}{keq\_FAFd} \\
& - 2 \cdot O2 (dissolved)\_lumen \cdot P700FA- \cdot kMehler\\
\frac{d B0}{dt} &= - B0 \cdot PPFD \cdot PSII\_cross\_section \\
& + B1 \cdot (kH0 + Q \cdot kH\_Qslope) + B1 \cdot kF \\
& + B2 \cdot Plastoquinone (oxidised) \cdot kPQred - \frac{B0 \cdot Plastoquinone (reduced) \cdot kPQred}{keq\_Plastoquinone (reduced)}\\
\frac{d B1}{dt} &= B0 \cdot PPFD \cdot PSII\_cross\_section \\
& - B1 \cdot (kH0 + Q \cdot kH\_Qslope) - B1 \cdot kF \\
& - B1 \cdot k2\\
\frac{d B2}{dt} &= B1 \cdot k2 \\
& - B2 \cdot Plastoquinone (oxidised) \cdot kPQred - \frac{B0 \cdot Plastoquinone (reduced) \cdot kPQred}{keq\_Plastoquinone (reduced)} \\
& - B2 \cdot PPFD \cdot PSII\_cross\_section + B3 \cdot kF \\
& + B3 \cdot (kH0 + Q \cdot kH\_Qslope)\\
\frac{d pH\_lumen}{dt} &= 0.04666666666666667 \cdot ATP\_pmf\_activity \cdot ATPactivity \cdot k\_ATPsynthase \cdot (\frac{ADP}{convf} - \frac{ATP}{convf \cdot kf\_atp\_synthase}) \\
& + 0.01 \cdot kf\_proton\_leak \cdot (protons\_lumen - 4000 \cdot {10}^{- pH}) \\
& - 0.01 \cdot B1 \cdot k2 \\
& - 0.04 \cdot \frac{Plastocyanine (oxidised) \cdot Plastoquinone (reduced) \cdot keq\_b6f\_dyn}{Plastoquinone (oxidised) + Plastoquinone (reduced)} - \frac{Plastocyanine (reduced) \cdot keq\_b6f\_dyn \cdot (1 - \frac{Plastoquinone (reduced)}{Plastoquinone (oxidised) + Plastoquinone (reduced)})}{keq\_b6f} \\
& + 0.01 \cdot \max(0, KEA3\_reg \cdot k\_KEA \cdot (K\_stroma \cdot protons\_lumen - K\_lumen \cdot protons)) \\
& - 0.04 \cdot \frac{0.9 \cdot Plastoquinone (oxidised) \cdot k\_NDH1 \cdot {10}^{-6.5 + pH\_lumen} \cdot {Ferredoxine (reduced)}^{2}}{(0.5 + {10}^{-6.5 + pH\_lumen}) \cdot (1 + 7.38905609893065 \cdot e^{- 100 \cdot P700+FA-})}\\
\frac{d pH}{dt} &= - 0.011666666666666667 \cdot ATP\_pmf\_activity \cdot ATPactivity \cdot k\_ATPsynthase \cdot (\frac{ADP}{convf} - \frac{ATP}{convf \cdot kf\_atp\_synthase}) \\
& - 0.0025 \cdot kf\_proton\_leak \cdot (protons\_lumen - 4000 \cdot {10}^{- pH}) \\
& + 0.0025 \cdot B2 \cdot Plastoquinone (oxidised) \cdot kPQred - \frac{B0 \cdot Plastoquinone (reduced) \cdot kPQred}{keq\_Plastoquinone (reduced)} \\
& + 0.01 \cdot \frac{Plastocyanine (oxidised) \cdot Plastoquinone (reduced) \cdot keq\_b6f\_dyn}{Plastoquinone (oxidised) + Plastoquinone (reduced)} - \frac{Plastocyanine (reduced) \cdot keq\_b6f\_dyn \cdot (1 - \frac{Plastoquinone (reduced)}{Plastoquinone (oxidised) + Plastoquinone (reduced)})}{keq\_b6f} \\
& - 0.0025 \cdot \max(0, KEA3\_reg \cdot k\_KEA \cdot (K\_stroma \cdot protons\_lumen - K\_lumen \cdot protons)) \\
& + 0.01 \cdot \frac{0.9 \cdot Plastoquinone (oxidised) \cdot k\_NDH1 \cdot {10}^{-6.5 + pH\_lumen} \cdot {Ferredoxine (reduced)}^{2}}{(0.5 + {10}^{-6.5 + pH\_lumen}) \cdot (1 + 7.38905609893065 \cdot e^{- 100 \cdot P700+FA-})}\\
\frac{d ATPactivity}{dt} &= \begin{cases}kActATPase \cdot (1 - ATPactivity) & PPFD > 0 \\ - ATPactivity \cdot kDeactATPase & \text{else}\end{cases}\\
\frac{d delta\_psi}{dt} &= - HPR \cdot volts\_per\_charge \cdot ATP\_pmf\_activity \cdot ATPactivity \cdot k\_ATPsynthase \cdot (\frac{ADP}{convf} - \frac{ATP}{convf \cdot kf\_atp\_synthase}) \\
& - 1 \cdot volts\_per\_charge \cdot kf\_proton\_leak \cdot (protons\_lumen - 4000 \cdot {10}^{- pH}) \\
& + volts\_per\_charge \cdot B1 \cdot k2 \\
& + 4 \cdot volts\_per\_charge \cdot \frac{Plastocyanine (oxidised) \cdot Plastoquinone (reduced) \cdot keq\_b6f\_dyn}{Plastoquinone (oxidised) + Plastoquinone (reduced)} - \frac{Plastocyanine (reduced) \cdot keq\_b6f\_dyn \cdot (1 - \frac{Plastoquinone (reduced)}{Plastoquinone (oxidised) + Plastoquinone (reduced)})}{keq\_b6f} \\
& - 1 \cdot volts\_per\_charge \cdot \frac{0.9 \cdot K\_lumen \cdot dG\_K\_ions \cdot perm\_K}{K\_stroma \cdot (1 + e^{1000 \cdot K\_delta\_psi\_treshold - 1000 \cdot delta\_psi})} \\
& - 1 \cdot volts\_per\_charge \cdot \frac{0.9 \cdot Cl\_driving\_force \cdot Cl\_stroma \cdot k\_VCCN1}{Cl\_lumen \cdot (1 + e^{1000 \cdot VCCN\_delta\_psi\_treshold - 1000 \cdot delta\_psi})} \\
& + volts\_per\_charge \cdot \frac{0.9 \cdot k\_Cl\_leak \cdot {Cl\_lumen - Cl\_stroma}^{2}}{total\_Cl\_2 \cdot (1 + e^{10 \cdot Cl\_leak\_PQ - 10 \cdot Plastoquinone (oxidised)})} \\
& + 4 \cdot volts\_per\_charge \cdot \frac{0.9 \cdot Plastoquinone (oxidised) \cdot k\_NDH1 \cdot {10}^{-6.5 + pH\_lumen} \cdot {Ferredoxine (reduced)}^{2}}{(0.5 + {10}^{-6.5 + pH\_lumen}) \cdot (1 + 7.38905609893065 \cdot e^{- 100 \cdot P700+FA-})}\\
\frac{d K\_stroma}{dt} &= - \max(0, KEA3\_reg \cdot k\_KEA \cdot (K\_stroma \cdot protons\_lumen - K\_lumen \cdot protons)) \\
& + \frac{0.9 \cdot K\_lumen \cdot dG\_K\_ions \cdot perm\_K}{K\_stroma \cdot (1 + e^{1000 \cdot K\_delta\_psi\_treshold - 1000 \cdot delta\_psi})}\\
\frac{d Cl\_stroma}{dt} &= - \frac{0.9 \cdot Cl\_driving\_force \cdot Cl\_stroma \cdot k\_VCCN1}{Cl\_lumen \cdot (1 + e^{1000 \cdot VCCN\_delta\_psi\_treshold - 1000 \cdot delta\_psi})} \\
& + \frac{0.9 \cdot k\_Cl\_leak \cdot {Cl\_lumen - Cl\_stroma}^{2}}{total\_Cl\_2 \cdot (1 + e^{10 \cdot Cl\_leak\_PQ - 10 \cdot Plastoquinone (oxidised)})} \\
& - ClCe\_activation \cdot k\_ClCe \cdot (Cl\_stroma - Cl\_lumen)
\end{align*}