Add analysis
Tomato KEA3 model
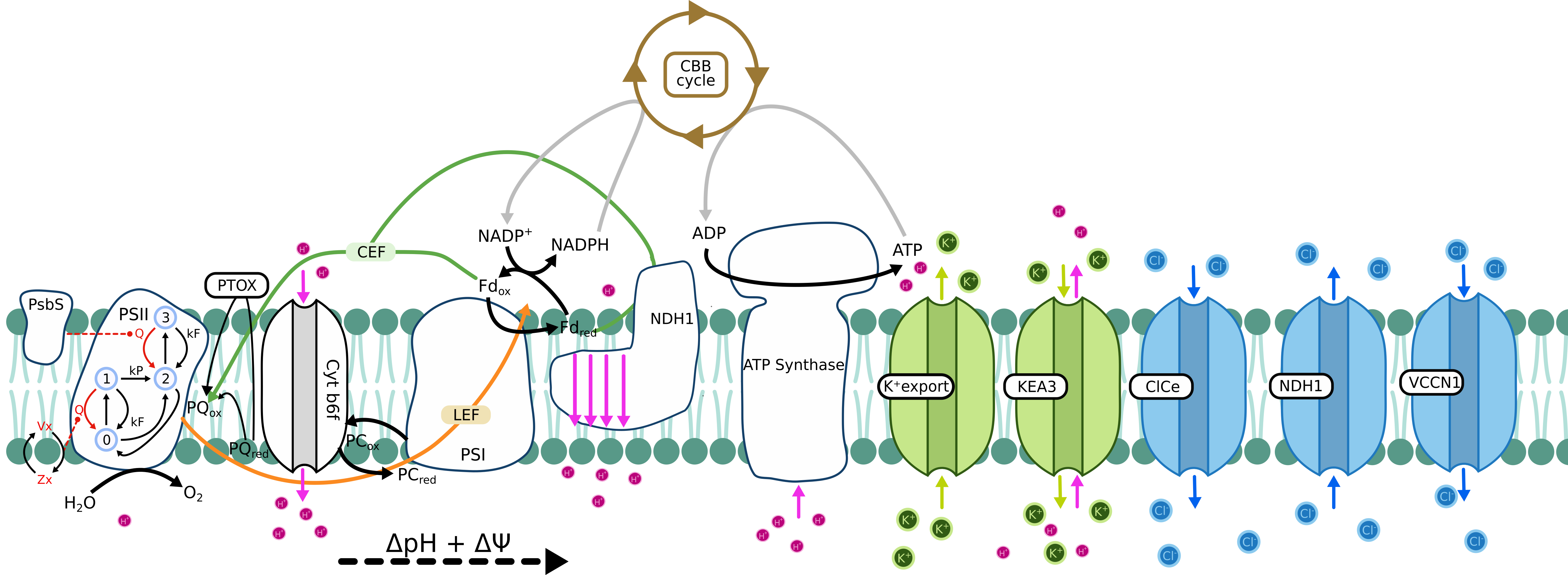
Simulation parameters
Analyses
Simulation - Normalised concentrations
Loading chart...
Simulation - Derived values
Loading chart...
PAM Fluorescence - NPQ results
Loading chart...
qL
Loading chart...
Model Details
Review and edit model structure, biological variables, and kinetic parameters.
| Name | Tex name | Initial value | Actions |
|---|---|---|---|
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Edit Options
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Eq Editor
Build an expression by selecting a node and replacing it with a MathML element, then adjust symbols to the allowed variable names.
Tip: click any element to select it, then choose a MathML element above or adjust its value.
Generated Python Code
import numpy as np
def model(
time: float,
variables: list[float],
):
B0, B1, B2, PQH2, ATP, H_lumen, delta_psi, Vx, PsbS, ATPactivity, K_lumen, K_stroma = variables
PPFD = 10
PSIItot = 2.5
PQtot = 20
APtot = 50
PsbStot = 1
Xtot = 1
O2ex = 8
Pi = 0.01
k_b6f = 0.22
pKreg = 6.4
kActATPase = 0.01
kDeactATPase = 0.002
kATPsynthase = 20
kATPconsumption = 10
HPR = 4.666666666666667
pKE0 = 7.211142552636095
b = 3.1924977471697407
kPQH2 = 250
kPTOX = 0.01
kH_Qslope = 5000000000
kH0 = 500000000
kF = 625000000
kP = 6939318750
pH_stroma = 7.8
kleak = 1000
bH = 100
kDeepoxV = 0.00096
kEpoxZ = 0.0013824
KphSatZ = 5.8
KZsat = 0.65
nHX = 5
nHZ = 3
nHL = 3
kDeprot = 0.0336
kProt = 0.07392
KphSatLHC = 5.8
gamma0 = 0.1
gamma1 = 1
gamma2 = 8
gamma3 = 2
pK_KEA3 = 6.75
k_KEA3 = 5
K_lumen_conc_initial = 0.1
K_stroma_conc_initial = 0.1
ATP_thres_KEA3 = 20.5
c = 0.1
F = 96.485
R = 0.0083
T = 298
E0QAQAm = -0.14
E0PQPQH2 = 0.354
E0PCPCm = 0.38
DeltaG0_ATP = 30.6
e = 2.71828
lumen_volume_per_area_membrane = 0.0014
stroma_volume_per_area_membrane = 0.0112
molChl_per_area_membrane = 0.00035
thylakoid_membrane_capacitance = 0.006
pHlumen_init = 7.2
RT = R * T
pH_lumen = - (np.log((0.001 * H_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane))) / (np.log(10))
H_lumen_conc = (0.001 * H_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane)
H_stroma = (1000 * stroma_volume_per_area_membrane * (10) ** (- pH_stroma)) / (molChl_per_area_membrane)
H_stroma_conc = (0.001 * H_stroma * molChl_per_area_membrane) / (stroma_volume_per_area_membrane)
delta_pH = pH_lumen + - pH_stroma
delta_pH_V = - (2.302585092994046 * R * T * delta_pH) / (F)
pmfV = delta_psi + - (2.302585092994046 * R * T * delta_pH) / (F)
volts_per_charge = (1 * F * molChl_per_area_membrane) / (thylakoid_membrane_capacitance)
PQ = PQtot + - PQH2
ADP = APtot + - ATP
PsbSP = PsbStot + - PsbS
Zx = Xtot + - Vx
Keq_PQH2 = np.exp((2 * E0PQPQH2 * F + - 2 * E0QAQAm * F + - 4.605170185988092 * R * T * pH_stroma) / (R * T))
Keqcytb6f = np.exp((- 2 * E0PQPQH2 * F + - 2 * F * pmfV + 2 * E0PCPCm * F + 4.605170185988092 * R * T * pH_lumen) / (R * T))
KeqATPsyn = Pi * np.exp((- DeltaG0_ATP + F * HPR * pmfV) / (R * T))
ATP_pmf_act = ((e) ** ((F * b * pmfV) / (R * T) + np.log((10) ** (- pKE0)))) / (1 + (e) ** ((F * b * pmfV) / (R * T) + np.log((10) ** (- pKE0))))
pHmod = 1 + - (1) / (1 + (10) ** (pH_lumen + - pKreg))
k_cytb6f = k_b6f * pHmod
Q0 = PsbS * Vx * gamma0
Q1 = PsbSP * Vx * gamma1
Q2 = PsbSP * Zx * gamma2
Q3 = PsbS * Zx * gamma3
Quencher_act = Q0 + Q1 + Q2 + Q3
B3 = PSIItot + - B0 + - B1 + - B2
rel_B0 = (B0) / (PSIItot)
rel_B1 = (B1) / (PSIItot)
rel_B2 = (B2) / (PSIItot)
rel_B3 = (B3) / (PSIItot)
qL = (B1 + B2) / (PSIItot)
Fluo = (B0 * kF) / (kF + kH0 + kP + Quencher_act * kH_Qslope) + (B2 * kF) / (kF + kH0 + Quencher_act * kH_Qslope)
PsbS_deprot_act = ((KZsat) ** (nHZ)) / ((KZsat) ** (nHZ) + (Zx) ** (nHZ))
K_stroma_conc = (0.001 * K_stroma * molChl_per_area_membrane) / (stroma_volume_per_area_membrane)
K_lumen_conc = (0.001 * K_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane)
reg_KEA3_ATP = (1 + - c) / (1 + np.exp((ATP + - ATP_thres_KEA3) / (c)))
reg_KEA3_pH = ((10) ** (pH_lumen + - pK_KEA3)) / (1 + (10) ** (pH_lumen + - pK_KEA3))
reg_KEA3 = reg_KEA3_ATP * reg_KEA3_pH
B01 = B0 * PPFD
B10Q = B1 * (kH0 + Quencher_act * kH_Qslope)
B10F = B1 * kF
vps2 = 0.5 * B1 * kP
B20 = B2 * PQ * kPQH2 + - (B0 * PQH2 * kPQH2) / (Keq_PQH2)
B23 = B2 * PPFD
B32F = B3 * kF
B32Q = B3 * (kH0 + Quencher_act * kH_Qslope)
vPQox = PQH2 * (O2ex * kPTOX + (Keqcytb6f * PPFD * k_cytb6f) / (1 + Keqcytb6f)) + - (PPFD * k_cytb6f * (PQtot + - PQH2)) / (1 + Keqcytb6f)
vATPactivity = kActATPase * (1 + - ATPactivity) if PPFD > 0 else - ATPactivity * kDeactATPase
vATPsynthase = ATP_pmf_act * ATPactivity * kATPsynthase * (ADP + - (ATP) / (KeqATPsyn))
vATPcons = ATP * kATPconsumption
vleak = kleak * (H_lumen_conc + - H_stroma_conc)
vXdeepox = (Vx * kDeepoxV * (H_lumen) ** (nHX)) / ((H_lumen) ** (nHX) + ((1000 * lumen_volume_per_area_membrane * (10) ** (- KphSatZ)) / (molChl_per_area_membrane)) ** (nHX))
vEpoxZ = Zx * kEpoxZ
vPsbSP = (PsbS * kProt * (H_lumen) ** (nHL)) / ((H_lumen) ** (nHL) + ((1000 * lumen_volume_per_area_membrane * (10) ** (- KphSatLHC)) / (molChl_per_area_membrane)) ** (nHL))
vPsbS = PsbSP * PsbS_deprot_act * kDeprot
vKEA3_in = max(0, (1000 * k_KEA3 * reg_KEA3 * stroma_volume_per_area_membrane * (H_lumen_conc * K_stroma_conc + - H_stroma_conc * K_lumen_conc)) / (molChl_per_area_membrane))
vKEA3_out = max(0, (1000 * k_KEA3 * lumen_volume_per_area_membrane * reg_KEA3 * (H_stroma_conc * K_lumen_conc + - H_lumen_conc * K_stroma_conc)) / (molChl_per_area_membrane))
dB0dt = -2*B01+2*B10Q+2*B10F+2*B20
dB1dt = +2*B01-2*B10Q-2*B10F-2*vps2
dB2dt = +2*vps2-2*B20-2*B23+2*B32F+2*B32Q
dPQH2dt = +B20-vPQox
dATPdt = +vATPsynthase-vATPcons
dH_lumendt = +((2) / (bH))*vps2+((4) / (bH))*vPQox+(- (HPR) / (bH))*vATPsynthase+(- (1) / (bH))*vleak+(- (1) / (bH))*vKEA3_in+((1) / (bH))*vKEA3_out
ddelta_psidt = +((2 * volts_per_charge) / (bH))*vps2+((4 * volts_per_charge) / (bH))*vPQox+(- (HPR * volts_per_charge) / (bH))*vATPsynthase+(- (volts_per_charge) / (bH))*vleak
dVxdt = -vXdeepox+vEpoxZ
dPsbSdt = -vPsbSP+vPsbS
dATPactivitydt = +vATPactivity
dK_lumendt = +vKEA3_in-vKEA3_out
dK_stromadt = -vKEA3_in+vKEA3_out
return [dB0dt, dB1dt, dB2dt, dPQH2dt, dATPdt, dH_lumendt, ddelta_psidt, dVxdt, dPsbSdt, dATPactivitydt, dK_lumendt, dK_stromadt]
def all_derived(
time: float,
variables: list[float],
):
B0, B1, B2, PQH2, ATP, H_lumen, delta_psi, Vx, PsbS, ATPactivity, K_lumen, K_stroma = variables
PPFD = 10
PSIItot = 2.5
PQtot = 20
APtot = 50
PsbStot = 1
Xtot = 1
O2ex = 8
Pi = 0.01
k_b6f = 0.22
pKreg = 6.4
kActATPase = 0.01
kDeactATPase = 0.002
kATPsynthase = 20
kATPconsumption = 10
HPR = 4.666666666666667
pKE0 = 7.211142552636095
b = 3.1924977471697407
kPQH2 = 250
kPTOX = 0.01
kH_Qslope = 5000000000
kH0 = 500000000
kF = 625000000
kP = 6939318750
pH_stroma = 7.8
kleak = 1000
bH = 100
kDeepoxV = 0.00096
kEpoxZ = 0.0013824
KphSatZ = 5.8
KZsat = 0.65
nHX = 5
nHZ = 3
nHL = 3
kDeprot = 0.0336
kProt = 0.07392
KphSatLHC = 5.8
gamma0 = 0.1
gamma1 = 1
gamma2 = 8
gamma3 = 2
pK_KEA3 = 6.75
k_KEA3 = 5
K_lumen_conc_initial = 0.1
K_stroma_conc_initial = 0.1
ATP_thres_KEA3 = 20.5
c = 0.1
F = 96.485
R = 0.0083
T = 298
E0QAQAm = -0.14
E0PQPQH2 = 0.354
E0PCPCm = 0.38
DeltaG0_ATP = 30.6
e = 2.71828
lumen_volume_per_area_membrane = 0.0014
stroma_volume_per_area_membrane = 0.0112
molChl_per_area_membrane = 0.00035
thylakoid_membrane_capacitance = 0.006
pHlumen_init = 7.2
RT = R * T
pH_lumen = - (np.log((0.001 * H_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane))) / (np.log(10))
H_lumen_conc = (0.001 * H_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane)
H_stroma = (1000 * stroma_volume_per_area_membrane * (10) ** (- pH_stroma)) / (molChl_per_area_membrane)
H_stroma_conc = (0.001 * H_stroma * molChl_per_area_membrane) / (stroma_volume_per_area_membrane)
delta_pH = pH_lumen + - pH_stroma
delta_pH_V = - (2.302585092994046 * R * T * delta_pH) / (F)
pmfV = delta_psi + - (2.302585092994046 * R * T * delta_pH) / (F)
volts_per_charge = (1 * F * molChl_per_area_membrane) / (thylakoid_membrane_capacitance)
PQ = PQtot + - PQH2
ADP = APtot + - ATP
PsbSP = PsbStot + - PsbS
Zx = Xtot + - Vx
Keq_PQH2 = np.exp((2 * E0PQPQH2 * F + - 2 * E0QAQAm * F + - 4.605170185988092 * R * T * pH_stroma) / (R * T))
Keqcytb6f = np.exp((- 2 * E0PQPQH2 * F + - 2 * F * pmfV + 2 * E0PCPCm * F + 4.605170185988092 * R * T * pH_lumen) / (R * T))
KeqATPsyn = Pi * np.exp((- DeltaG0_ATP + F * HPR * pmfV) / (R * T))
ATP_pmf_act = ((e) ** ((F * b * pmfV) / (R * T) + np.log((10) ** (- pKE0)))) / (1 + (e) ** ((F * b * pmfV) / (R * T) + np.log((10) ** (- pKE0))))
pHmod = 1 + - (1) / (1 + (10) ** (pH_lumen + - pKreg))
k_cytb6f = k_b6f * pHmod
Q0 = PsbS * Vx * gamma0
Q1 = PsbSP * Vx * gamma1
Q2 = PsbSP * Zx * gamma2
Q3 = PsbS * Zx * gamma3
Quencher_act = Q0 + Q1 + Q2 + Q3
B3 = PSIItot + - B0 + - B1 + - B2
rel_B0 = (B0) / (PSIItot)
rel_B1 = (B1) / (PSIItot)
rel_B2 = (B2) / (PSIItot)
rel_B3 = (B3) / (PSIItot)
qL = (B1 + B2) / (PSIItot)
Fluo = (B0 * kF) / (kF + kH0 + kP + Quencher_act * kH_Qslope) + (B2 * kF) / (kF + kH0 + Quencher_act * kH_Qslope)
PsbS_deprot_act = ((KZsat) ** (nHZ)) / ((KZsat) ** (nHZ) + (Zx) ** (nHZ))
K_stroma_conc = (0.001 * K_stroma * molChl_per_area_membrane) / (stroma_volume_per_area_membrane)
K_lumen_conc = (0.001 * K_lumen * molChl_per_area_membrane) / (lumen_volume_per_area_membrane)
reg_KEA3_ATP = (1 + - c) / (1 + np.exp((ATP + - ATP_thres_KEA3) / (c)))
reg_KEA3_pH = ((10) ** (pH_lumen + - pK_KEA3)) / (1 + (10) ** (pH_lumen + - pK_KEA3))
reg_KEA3 = reg_KEA3_ATP * reg_KEA3_pH
B01 = B0 * PPFD
B10Q = B1 * (kH0 + Quencher_act * kH_Qslope)
B10F = B1 * kF
vps2 = 0.5 * B1 * kP
B20 = B2 * PQ * kPQH2 + - (B0 * PQH2 * kPQH2) / (Keq_PQH2)
B23 = B2 * PPFD
B32F = B3 * kF
B32Q = B3 * (kH0 + Quencher_act * kH_Qslope)
vPQox = PQH2 * (O2ex * kPTOX + (Keqcytb6f * PPFD * k_cytb6f) / (1 + Keqcytb6f)) + - (PPFD * k_cytb6f * (PQtot + - PQH2)) / (1 + Keqcytb6f)
vATPactivity = kActATPase * (1 + - ATPactivity) if PPFD > 0 else - ATPactivity * kDeactATPase
vATPsynthase = ATP_pmf_act * ATPactivity * kATPsynthase * (ADP + - (ATP) / (KeqATPsyn))
vATPcons = ATP * kATPconsumption
vleak = kleak * (H_lumen_conc + - H_stroma_conc)
vXdeepox = (Vx * kDeepoxV * (H_lumen) ** (nHX)) / ((H_lumen) ** (nHX) + ((1000 * lumen_volume_per_area_membrane * (10) ** (- KphSatZ)) / (molChl_per_area_membrane)) ** (nHX))
vEpoxZ = Zx * kEpoxZ
vPsbSP = (PsbS * kProt * (H_lumen) ** (nHL)) / ((H_lumen) ** (nHL) + ((1000 * lumen_volume_per_area_membrane * (10) ** (- KphSatLHC)) / (molChl_per_area_membrane)) ** (nHL))
vPsbS = PsbSP * PsbS_deprot_act * kDeprot
vKEA3_in = max(0, (1000 * k_KEA3 * reg_KEA3 * stroma_volume_per_area_membrane * (H_lumen_conc * K_stroma_conc + - H_stroma_conc * K_lumen_conc)) / (molChl_per_area_membrane))
vKEA3_out = max(0, (1000 * k_KEA3 * lumen_volume_per_area_membrane * reg_KEA3 * (H_stroma_conc * K_lumen_conc + - H_lumen_conc * K_stroma_conc)) / (molChl_per_area_membrane))
return [RT, pH_lumen, H_lumen_conc, H_stroma, H_stroma_conc, delta_pH, delta_pH_V, pmfV, volts_per_charge, PQ, ADP, PsbSP, Zx, Keq_PQH2, Keqcytb6f, KeqATPsyn, ATP_pmf_act, pHmod, k_cytb6f, Q0, Q1, Q2, Q3, Quencher_act, B3, rel_B0, rel_B1, rel_B2, rel_B3, qL, Fluo, PsbS_deprot_act, K_stroma_conc, K_lumen_conc, reg_KEA3_ATP, reg_KEA3_pH, reg_KEA3, B01, B10Q, B10F, vps2, B20, B23, B32F, B32Q, vPQox, vATPactivity, vATPsynthase, vATPcons, vleak, vXdeepox, vEpoxZ, vPsbSP, vPsbS, vKEA3_in, vKEA3_out]
derived = all_derived
y0 = {"B0": 2.5, "B1": 0, "B2": 0, "PQH2": 0, "ATP": 25, "H_lumen": 0.00025238293779207717, "delta_psi": 0, "Vx": 1, "PsbS": 1, "ATPactivity": 0.1, "K_lumen": 399.99999999999994, "K_stroma": 3199.9999999999995}
Generated LaTeX Code
\begin{align*}
\frac{d B0}{dt} &= - 2 \cdot B0 \cdot PPFD \\
& + 2 \cdot B1 \cdot (kH0 + Quencher act. \cdot kH\_Qslope) \\
& + 2 \cdot B1 \cdot kF \\
& + 2 \cdot B2 \cdot PQ \cdot kPQH2 - \frac{B0 \cdot PQH2 \cdot kPQH2}{Keq\_PQH2}\\
\frac{d B1}{dt} &= 2 \cdot B0 \cdot PPFD \\
& - 2 \cdot B1 \cdot (kH0 + Quencher act. \cdot kH\_Qslope) \\
& - 2 \cdot B1 \cdot kF - 2 \cdot 0.5 \cdot B1 \cdot kP\\
\frac{d B2}{dt} &= 2 \cdot 0.5 \cdot B1 \cdot kP \\
& - 2 \cdot B2 \cdot PQ \cdot kPQH2 - \frac{B0 \cdot PQH2 \cdot kPQH2}{Keq\_PQH2} \\
& - 2 \cdot B2 \cdot PPFD + 2 \cdot B3 \cdot kF \\
& + 2 \cdot B3 \cdot (kH0 + Quencher act. \cdot kH\_Qslope)\\
\frac{d PQH2}{dt} &= B2 \cdot PQ \cdot kPQH2 - \frac{B0 \cdot PQH2 \cdot kPQH2}{Keq\_PQH2} \\
& - PQH2 \cdot (O2ex \cdot kPTOX + \frac{Keqcytb6f \cdot PPFD \cdot k\_cytb6f}{1 + Keqcytb6f}) - \frac{PPFD \cdot k\_cytb6f \cdot (PQtot - PQH2)}{1 + Keqcytb6f}\\
\frac{d ATP}{dt} &= ATP\_pmf\_act \cdot ATPactivity \cdot kATPsynthase \cdot (ADP - \frac{ATP}{KeqATPsyn}) \\
& - ATP \cdot kATPconsumption\\
\frac{d H\_lumen}{dt} &= \frac{2}{bH} \cdot 0.5 \cdot B1 \cdot kP \\
& + \frac{4}{bH} \cdot PQH2 \cdot (O2ex \cdot kPTOX + \frac{Keqcytb6f \cdot PPFD \cdot k\_cytb6f}{1 + Keqcytb6f}) - \frac{PPFD \cdot k\_cytb6f \cdot (PQtot - PQH2)}{1 + Keqcytb6f} \\
& - \frac{HPR}{bH} \cdot ATP\_pmf\_act \cdot ATPactivity \cdot kATPsynthase \cdot (ADP - \frac{ATP}{KeqATPsyn}) \\
& - \frac{1}{bH} \cdot kleak \cdot (H\_lumen\_conc - H\_stroma\_conc) \\
& - \frac{1}{bH} \cdot \max(0, \frac{1000 \cdot k\_KEA3 \cdot reg\_KEA3 \cdot stroma\_volume\_per\_area\_membrane \cdot (H\_lumen\_conc \cdot K\_stroma\_conc - H\_stroma\_conc \cdot K\_lumen\_conc)}{molChl\_per\_area\_membrane}) \\
& + \frac{1}{bH} \cdot \max(0, \frac{1000 \cdot k\_KEA3 \cdot lumen\_volume\_per\_area\_membrane \cdot reg\_KEA3 \cdot (H\_stroma\_conc \cdot K\_lumen\_conc - H\_lumen\_conc \cdot K\_stroma\_conc)}{molChl\_per\_area\_membrane})\\
\frac{d delta\_psi}{dt} &= \frac{2 \cdot volts\_per\_charge}{bH} \cdot 0.5 \cdot B1 \cdot kP \\
& + \frac{4 \cdot volts\_per\_charge}{bH} \cdot PQH2 \cdot (O2ex \cdot kPTOX + \frac{Keqcytb6f \cdot PPFD \cdot k\_cytb6f}{1 + Keqcytb6f}) - \frac{PPFD \cdot k\_cytb6f \cdot (PQtot - PQH2)}{1 + Keqcytb6f} \\
& - \frac{HPR \cdot volts\_per\_charge}{bH} \cdot ATP\_pmf\_act \cdot ATPactivity \cdot kATPsynthase \cdot (ADP - \frac{ATP}{KeqATPsyn}) \\
& - \frac{volts\_per\_charge}{bH} \cdot kleak \cdot (H\_lumen\_conc - H\_stroma\_conc)\\
\frac{d Vx}{dt} &= - \frac{Vx \cdot kDeepoxV \cdot {H\_lumen}^{nHX}}{{H\_lumen}^{nHX} + {\frac{1000 \cdot lumen\_volume\_per\_area\_membrane \cdot {10}^{- KphSatZ}}{molChl\_per\_area\_membrane}}^{nHX}} \\
& + Zx \cdot kEpoxZ\\
\frac{d PsbS}{dt} &= - \frac{PsbS \cdot kProt \cdot {H\_lumen}^{nHL}}{{H\_lumen}^{nHL} + {\frac{1000 \cdot lumen\_volume\_per\_area\_membrane \cdot {10}^{- KphSatLHC}}{molChl\_per\_area\_membrane}}^{nHL}} \\
& + PsbSP \cdot PsbS\_deprot\_act \cdot kDeprot\\
\frac{d ATPactivity}{dt} &= \begin{cases}kActATPase \cdot (1 - ATPactivity) & PPFD > 0 \\ - ATPactivity \cdot kDeactATPase & \text{else}\end{cases}\\
\frac{d K\_lumen}{dt} &= \max(0, \frac{1000 \cdot k\_KEA3 \cdot reg\_KEA3 \cdot stroma\_volume\_per\_area\_membrane \cdot (H\_lumen\_conc \cdot K\_stroma\_conc - H\_stroma\_conc \cdot K\_lumen\_conc)}{molChl\_per\_area\_membrane}) \\
& - \max(0, \frac{1000 \cdot k\_KEA3 \cdot lumen\_volume\_per\_area\_membrane \cdot reg\_KEA3 \cdot (H\_stroma\_conc \cdot K\_lumen\_conc - H\_lumen\_conc \cdot K\_stroma\_conc)}{molChl\_per\_area\_membrane})\\
\frac{d K\_stroma}{dt} &= - \max(0, \frac{1000 \cdot k\_KEA3 \cdot reg\_KEA3 \cdot stroma\_volume\_per\_area\_membrane \cdot (H\_lumen\_conc \cdot K\_stroma\_conc - H\_stroma\_conc \cdot K\_lumen\_conc)}{molChl\_per\_area\_membrane}) \\
& + \max(0, \frac{1000 \cdot k\_KEA3 \cdot lumen\_volume\_per\_area\_membrane \cdot reg\_KEA3 \cdot (H\_stroma\_conc \cdot K\_lumen\_conc - H\_lumen\_conc \cdot K\_stroma\_conc)}{molChl\_per\_area\_membrane})
\end{align*}