from dataclasses import dataclass
from mxlpy import Model, Simulator, fns, plot
from mxlpy.types import InitialAssignment
def one_div(x: float) -> float:
return 1.0 / x
def minus_one_div(x: float) -> float:
return -1.0 / x
def times_frac(top: float, btm: float, x: float) -> float:
return top / btm * x
def ma_1s_1p_keq(s1: float, p1: float, kf: float, keq: float) -> float:
return kf * (s1 - p1 / keq)
Compartmental models¶
We follow the recommendations by Hofmeyr (2020) (https://doi.org/10.1016/j.biosystems.2020.104203) when it comes to compartmentalisation. That is, describe compartmentalised variables in terms of amounts instead of concentration and scale kinetic constants by the compartment to yield rate factors.
Depending on your use case, your experimental data might be described as concentrations and communication is also prefered in concentrations.
mxlpy provides various ways of how you can automate that mapping
- use
Derivedvariables and parameters to obtain concentrations from the amounts. - use
InitialAssignments to calculate initial amounts in case your data is given in concentrations
Below we define convenience functions for exactly that, to reduce boilerplate code
@dataclass
class VarNames:
amount: str
conc: str
def add_cvar(
model: Model,
name: str,
compartment: str,
initial: float,
*,
initial_is_amount: bool = True,
amount_prefix: str = "n_",
conc_prefix: str = "c_",
) -> VarNames:
amount = f"{amount_prefix}{name}_{compartment}"
if not initial_is_amount:
# FIXME: rewrite initial assignment to take in float?
model.add_parameter(init_c := f"c_{name}_init", initial)
model.add_variable(
amount,
InitialAssignment(fn=fns.mul, args=[init_c, compartment]),
)
else:
model.add_variable(amount, initial)
model.add_derived(
conc := f"{conc_prefix}{name}_{compartment}",
fn=fns.div,
args=[amount, compartment],
)
return VarNames(amount, conc)
m = Model()
m.add_parameters({"c1": 2.0, "c2": 4.0})
# Add a compartmentalised variable with an initial amount
x_c1 = add_cvar(m, "x", compartment="c1", initial=1.5)
# Add a compartmentalised variable with an initial concentration
x_c2 = add_cvar(m, "x", compartment="c2", initial=1.5, initial_is_amount=False)
args = m.get_args()
print("Amounts", args.loc[[x_c1.amount, x_c2.amount]], sep="\n", end="\n\n")
print("Concentrations", args.loc[[x_c1.conc, x_c2.conc]], sep="\n", end="\n\n")
Amounts n_x_c1 1.5 n_x_c2 6.0 dtype: float64 Concentrations c_x_c1 0.75 c_x_c2 1.50 dtype: float64
Here is another quick conveniece function to quickly kinetic constants and their respective **rate factors **
def rate_factor(rxn_place: float, k: float, compartment: float) -> float:
return rxn_place * k / compartment
def add_rate_factor(
model: Model,
base: str,
value: float,
rxn_compartment: str,
cpd_compartment: str,
k_prefix: str = "k_",
f_prefix: str = "f_",
) -> str:
model.add_parameter(k := f"{k_prefix}{base}", value)
f = f"{f_prefix}{base}"
model.add_derived(
f,
fn=rate_factor,
args=[rxn_compartment, k, cpd_compartment],
)
return f
m = Model()
m.add_parameters({"c1": 2.0, "a1": 4.0})
rf = add_rate_factor(m, "r1", 1.0, rxn_compartment="a1", cpd_compartment="c1")
m.get_args().loc[rf]
np.float64(2.0)
Examples¶
Single compartment¶
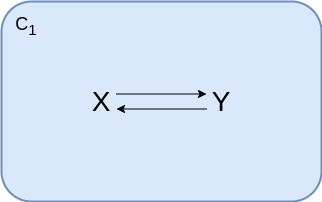
In case all your reactions happen in the volume of the same compartment, you don't need to change anything.
Whether your variables describe an amount or a concentration does not matter, the equations are identical.
Here we use standard reversible mass-action kinetics, once formulated explicitly with forward and backward kinetic constants and once using the equilibrium constant.
$$\begin{align*} v &= k_f x - k_r y \\ &= k_f \left( x - \frac{y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"x": 2.0, "y": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0})
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["x", "y", "kf", "keq"],
stoichiometry={"x": -1, "y": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
_ = plot.lines(
c,
xlabel="Time",
ylabel="Amount or Concentration",
ax=plot.one_axes(figsize=(4, 3))[1],
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[4], line 13 9 stoichiometry={"x": -1, "y": 1}, 10 ) 11 ) 12 ---> 13 c, v = unwrap(Simulator(m).simulate(10).get_result()) 14 _ = plot.lines( 15 c, 16 xlabel="Time", NameError: name 'unwrap' is not defined
Single compartment - at membrane¶

$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{cyan}C_1} } - k_r \frac{n_y}{{\color{cyan}C_1}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"n_x": 2.0, "n_y": 1.0})
.add_parameters(
{
"kf": 1.0,
"keq": 2.0,
"c1": 1.5, # compartment volume
"a1": 1.0, # area of membrane
}
)
# Now compartmentalise
.add_derived("ff", rate_factor, args=["a1", "kf", "c1"])
.add_derived("x_c1", fns.div, args=["n_x", "c1"])
.add_derived("y_c1", fns.div, args=["n_y", "c1"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["n_x", "n_y", "ff", "keq"],
stoichiometry={"n_x": -1, "n_y": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["n_x", "n_y"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1", "y_c1"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[5], line 24 20 stoichiometry={"n_x": -1, "n_y": 1}, 21 ) 22 ) 23 ---> 24 c, v = unwrap(Simulator(m).simulate(10).get_result()) 25 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 26 _ = plot.lines( 27 c.loc[:, ["n_x", "n_y"]], NameError: name 'unwrap' is not defined
Single compartment - on membrane¶

This special case depends only on the area of the membrane, not on the volume of the compartment
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{purple} A} } - k_r \frac{n_y}{{\color{purple} A} } \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"n_x": 2.0, "n_y": 1.0})
.add_parameters(
{
"kf": 1.0,
"keq": 2.0,
}
)
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["n_x", "n_y", "kf", "keq"],
stoichiometry={"n_x": -1, "n_y": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
_ = plot.lines(
c.loc[:, ["n_x", "n_y"]],
xlabel="Time",
ylabel="Amount",
ax=plot.one_axes(figsize=(4, 3))[1],
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[6], line 18 14 stoichiometry={"n_x": -1, "n_y": 1}, 15 ) 16 ) 17 ---> 18 c, v = unwrap(Simulator(m).simulate(10).get_result()) 19 _ = plot.lines( 20 c.loc[:, ["n_x", "n_y"]], 21 xlabel="Time", NameError: name 'unwrap' is not defined
Two compartments - diffusion accross membrane¶
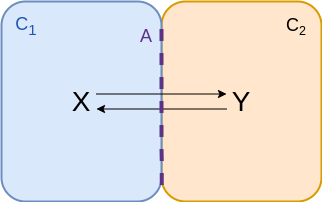
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{cyan} C_1}} - k_r \frac{n_y}{{\color{coral} C_2}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_c1": 2.0, "ny_c2": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c1": 1.5, "c2": 0.5})
# Now compartmentalise
.add_derived("ff", fns.div, args=["kf", "c1"])
.add_derived("x_c1", fns.div, args=["nx_c1", "c1"])
.add_derived("x_c2", fns.div, args=["ny_c2", "c2"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_c1", "ny_c2", "ff", "keq"],
stoichiometry={"nx_c1": -1, "ny_c2": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_c1", "ny_c2"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1", "x_c2"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[7], line 17 13 stoichiometry={"nx_c1": -1, "ny_c2": 1}, 14 ) 15 ) 16 ---> 17 c, v = unwrap(Simulator(m).simulate(10).get_result()) 18 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 19 _ = plot.lines( 20 c.loc[:, ["nx_c1", "ny_c2"]], NameError: name 'unwrap' is not defined
Two compartments - inside¶
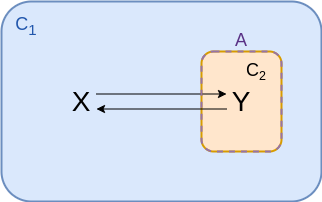
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{cyan} C_1}} - k_r \frac{n_y}{{\color{coral} C_2}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_c1": 2.0, "ny_c2": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c1": 1.5, "c2": 0.5})
# Now compartmentalise
.add_derived("ff", fns.div, args=["kf", "c1"])
.add_derived("x_c1", fns.div, args=["nx_c1", "c1"])
.add_derived("x_c2", fns.div, args=["ny_c2", "c2"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_c1", "ny_c2", "ff", "keq"],
stoichiometry={"nx_c1": -1, "ny_c2": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_c1", "ny_c2"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1", "x_c2"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[8], line 17 13 stoichiometry={"nx_c1": -1, "ny_c2": 1}, 14 ) 15 ) 16 ---> 17 c, v = unwrap(Simulator(m).simulate(10).get_result()) 18 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 19 _ = plot.lines( 20 c.loc[:, ["nx_c1", "ny_c2"]], NameError: name 'unwrap' is not defined
Two compartments - at membrane¶
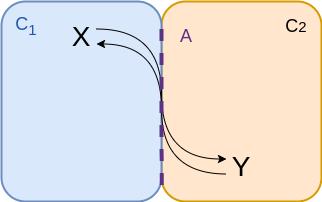
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{cyan} C_1}} - k_r \frac{n_y}{{\color{coral} C_2}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_c1": 2.0, "ny_c2": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c1": 1.5, "c2": 0.5})
# Now compartmentalise
.add_derived("ff", fns.div, args=["kf", "c1"])
.add_derived("x_c1", fns.div, args=["nx_c1", "c1"])
.add_derived("x_c2", fns.div, args=["ny_c2", "c2"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_c1", "ny_c2", "ff", "keq"],
stoichiometry={"nx_c1": -1, "ny_c2": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_c1", "ny_c2"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1", "x_c2"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[9], line 17 13 stoichiometry={"nx_c1": -1, "ny_c2": 1}, 14 ) 15 ) 16 ---> 17 c, v = unwrap(Simulator(m).simulate(10).get_result()) 18 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 19 _ = plot.lines( 20 c.loc[:, ["nx_c1", "ny_c2"]], NameError: name 'unwrap' is not defined
Two compartments - on membrane¶
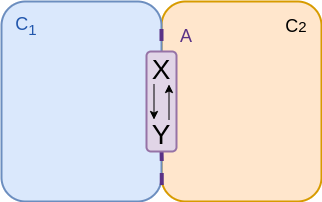
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{purple} A}} - k_r \frac{n_y}{{\color{purple} A}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_c1": 2.0, "ny_c2": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c1": 1.5, "c2": 0.5})
# Now compartmentalise
.add_derived("ff", fns.div, args=["kf", "c1"])
.add_derived("x_c1", fns.div, args=["nx_c1", "c1"])
.add_derived("x_c2", fns.div, args=["ny_c2", "c2"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_c1", "ny_c2", "ff", "keq"],
stoichiometry={"nx_c1": -1, "ny_c2": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_c1", "ny_c2"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1", "x_c2"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[10], line 17 13 stoichiometry={"nx_c1": -1, "ny_c2": 1}, 14 ) 15 ) 16 ---> 17 c, v = unwrap(Simulator(m).simulate(10).get_result()) 18 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 19 _ = plot.lines( 20 c.loc[:, ["nx_c1", "ny_c2"]], NameError: name 'unwrap' is not defined
Two compartments - onto / off membrane¶
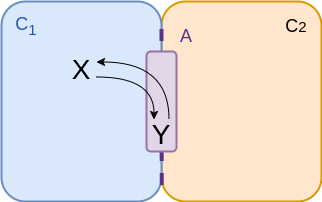
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{cyan} C_1}} - k_r \frac{n_y}{{\color{purple} A} } \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_c1": 2.0, "ny_a": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c1": 1.5, "a": 0.5})
# Now compartmentalise
.add_derived("ff", rate_factor, args=["a", "kf", "c1"])
.add_derived("x_c1", fns.div, args=["nx_c1", "c1"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_c1", "ny_a", "ff", "keq"],
stoichiometry={"nx_c1": -1, "ny_a": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_c1", "ny_a"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["x_c1"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[11], line 16 12 stoichiometry={"nx_c1": -1, "ny_a": 1}, 13 ) 14 ) 15 ---> 16 c, v = unwrap(Simulator(m).simulate(10).get_result()) 17 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 18 _ = plot.lines( 19 c.loc[:, ["nx_c1", "ny_a"]], NameError: name 'unwrap' is not defined
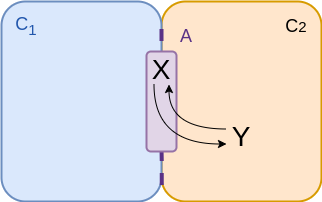
$$\begin{align*} v &= {\color{purple} A} \left( k_f \frac{n_x}{{\color{purple} A} } - k_r \frac{n_y}{{\color{coral} C_2}} \right) \\ &= f_f n_x - f_r n_y \\ &= f_f \left( n_x - \frac{n_y}{K_{eq}} \right)\\ \end{align*}$$
m = (
Model()
.add_variables({"nx_a": 2.0, "ny_c2": 1.0})
.add_parameters({"kf": 1.0, "keq": 2.0, "c2": 1.5, "a": 0.5})
# Now compartmentalise
.add_derived("ff", rate_factor, args=["a", "kf", "a"])
.add_derived("y_c2", fns.div, args=["nx_a", "c2"])
.add_reaction(
"v1",
ma_1s_1p_keq,
args=["nx_a", "ny_c2", "ff", "keq"],
stoichiometry={"nx_a": -1, "ny_c2": 1},
)
)
c, v = unwrap(Simulator(m).simulate(10).get_result())
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3))
_ = plot.lines(
c.loc[:, ["nx_a", "ny_c2"]],
ax=ax1,
xlabel="Time",
ylabel="Amount",
)
_ = plot.lines(
c.loc[:, ["y_c2"]],
ax=ax2,
xlabel="Time",
ylabel="Concentration",
)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[12], line 16 12 stoichiometry={"nx_a": -1, "ny_c2": 1}, 13 ) 14 ) 15 ---> 16 c, v = unwrap(Simulator(m).simulate(10).get_result()) 17 fig, (ax1, ax2) = plot.two_axes(figsize=(6, 3)) 18 _ = plot.lines( 19 c.loc[:, ["nx_a", "ny_c2"]], NameError: name 'unwrap' is not defined
First finish line
With that you now know most of what you will need from a day-to-day basis about compartmentalisation in mxlpy.Congratulations!