from __future__ import annotations
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import mxlpy as mb2
from example_models import (
get_lin_chain_two_circles,
get_linear_chain_2v,
get_upper_glycolysis,
)
from mxlpy import make_protocol, mc, mca, plot
Monte Carlo methods¶
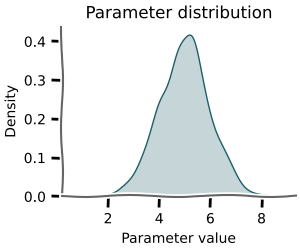
Almost every parameter in biology is better described with a distribution than a single value.
Monte-carlo methods allow you to capture the range of possible behaviour your model can exhibit.
This is especially useful when you want to understand the uncertainty in your model's predictions.
mxlpy offers these Monte Carlo methods for all scans ...
 +
+
 =
=
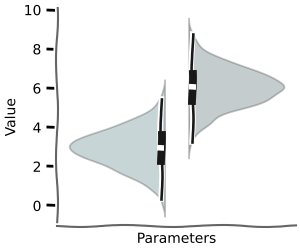
and even for metabolic control analysis
 +
+
 =
=
In this tutorial we will mostly use the mxlpy.distributions and mxlpy.mca modules, which contain the functionality to sample from distributions and run distributed analyses.
Sample values¶
To do any Monte-Carlo analysis, we first need to be able to sample values.
For that, you can use the sample function and distributions supplied by mxlpy.
These are mostly thin wrappers around the numpy and scipy sampling methods.
from mxlpy.distributions import LogNormal, Uniform, sample
sample(
{
"k2": Uniform(1.0, 2.0),
"k3": LogNormal(mean=1.0, sigma=1.0),
},
n=5,
)
| k2 | k3 | |
|---|---|---|
| 0 | 1.773956 | 0.739205 |
| 1 | 1.438878 | 3.088978 |
| 2 | 1.858598 | 1.981308 |
| 3 | 1.697368 | 2.672993 |
| 4 | 1.094177 | 1.158303 |
Steady-state¶
Using mc.steady_state you can calculate the steady-state distribution given the monte-carlo parameters.
This works analogously to the scan.steady_state function, except the index of the dataframes is always just an integer.
The parameters used can be obtained by result.parameters.
We will use a linear chain of reactions with two circles as an example model for this notebook.
$$ \begin{array}{c|c} \mathrm{Reaction} & \mathrm{Stoichiometry} \\ \hline v_0 & \varnothing \rightarrow{} \mathrm{x_1} \\ v_1 & -\mathrm{x_1} \rightarrow{} \mathrm{x_2} \\ v_2 & -\mathrm{x_1} \rightarrow{} \mathrm{x_3} \\ v_3 & -\mathrm{x_1} \rightarrow{} \mathrm{x_4} \\ v_4 & -\mathrm{x_4} \rightarrow{} \varnothing\\ v_5 & -\mathrm{x_2} \rightarrow{} \mathrm{x_1} \\ v_6 & -\mathrm{x_3} \rightarrow{} \mathrm{x_1} \\ \end{array} $$
ss = mc.steady_state(
get_linear_chain_2v(),
mc_to_scan=sample(
{
"k1": Uniform(0.9, 1.1),
"k2": Uniform(1.0, 1.3),
"k3": LogNormal(mean=1.0, sigma=0.2),
},
n=10,
),
)
fig, (ax1, ax2) = plot.two_axes(figsize=(6, 2.5), sharex=False)
plot.violins(ss.variables, ax=ax1)
plot.violins(ss.fluxes, ax=ax2)
ax1.set(xlabel="Variables", ylabel="Concentration / a.u.")
ax2.set(xlabel="Reactions", ylabel="Flux / a.u.")
plt.show()
0%| | 0/10 [00:00<?, ?it/s]
10%|█ | 1/10 [00:05<00:45, 5.10s/it]
30%|███ | 3/10 [00:05<00:09, 1.38s/it]
100%|██████████| 10/10 [00:05<00:00, 3.29it/s]
100%|██████████| 10/10 [00:05<00:00, 1.87it/s]
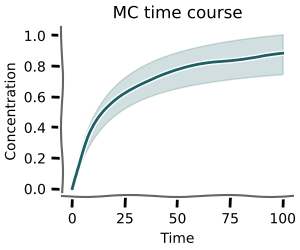
Time course¶
Using mc.time_course you can calculate time courses for sampled parameters.
 +
+
 =
=

This function works analogously to scan.time_course.
The pandas.DataFrames for concentrations and fluxes have a n x time pandas.MultiIndex.
The corresponding parameters can be found in result.parameters
tc = mc.time_course(
get_linear_chain_2v(),
time_points=np.linspace(0, 1, 11),
mc_to_scan=sample(
{
"k1": Uniform(0.9, 1.1),
"k2": Uniform(1.0, 1.3),
"k3": LogNormal(mean=1.0, sigma=0.2),
},
n=10,
),
)
fig, (ax1, ax2) = plot.two_axes(figsize=(7, 4))
plot.lines_mean_std_from_2d_idx(tc.variables, ax=ax1)
plot.lines_mean_std_from_2d_idx(tc.fluxes, ax=ax2)
ax1.set(xlabel="Time / a.u", ylabel="Concentration / a.u.")
ax2.set(xlabel="Time / a.u", ylabel="Flux / a.u.")
plt.show()
0%| | 0/10 [00:00<?, ?it/s]
10%|█ | 1/10 [00:04<00:44, 4.98s/it]
50%|█████ | 5/10 [00:05<00:03, 1.28it/s]
100%|██████████| 10/10 [00:05<00:00, 1.93it/s]
Protocol time course¶
Using mc.time_course_over_protocol you can calculate time courses for sampled parameters given a discrete protocol.
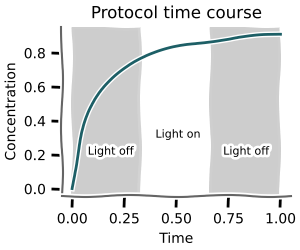 +
+
 =
=
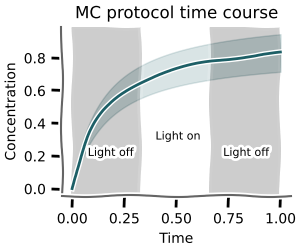
The pandas.DataFrames for concentrations and fluxes have a n x time pandas.MultiIndex.
The corresponding parameters can be found in scan.parameters
tc = mc.protocol_time_course(
get_linear_chain_2v(),
time_points=np.linspace(0, 6, 21),
protocol=make_protocol(
[
(1, {"k1": 1}),
(2, {"k1": 2}),
(3, {"k1": 1}),
]
),
mc_to_scan=sample(
{
"k2": Uniform(1.0, 1.3),
"k3": LogNormal(mean=1.0, sigma=0.2),
},
n=10,
),
)
fig, (ax1, ax2) = plot.two_axes(figsize=(7, 4))
plot.lines_mean_std_from_2d_idx(tc.variables, ax=ax1)
plot.lines_mean_std_from_2d_idx(tc.fluxes, ax=ax2)
for ax in (ax1, ax2):
plot.shade_protocol(tc.protocol["k1"], ax=ax, alpha=0.1)
ax1.set(xlabel="Time / a.u", ylabel="Concentration / a.u.")
ax2.set(xlabel="Time / a.u", ylabel="Flux / a.u.")
plt.show()
0%| | 0/10 [00:00<?, ?it/s]
0%| | 0/10 [00:04<?, ?it/s]
--------------------------------------------------------------------------- _RemoteTraceback Traceback (most recent call last) _RemoteTraceback: """ Traceback (most recent call last): File "/home/runner/work/MxlPy/MxlPy/.venv/lib/python3.13/site-packages/loky/process_executor.py", line 490, in _process_worker r = call_item() File "/home/runner/work/MxlPy/MxlPy/.venv/lib/python3.13/site-packages/loky/process_executor.py", line 291, in __call__ return self.fn(*self.args, **self.kwargs) ~~~~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^ File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/parallel.py", line 94, in _load_or_run res = fn(v) File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/scan.py", line 80, in _update_parameters_and_initial_conditions return fn(model) File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/scan.py", line 315, in _protocol_time_course_worker .simulate_protocol_time_course( ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^ protocol=protocol, ^^^^^^^^^^^^^^^^^^ time_points=time_points, ^^^^^^^^^^^^^^^^^^^^^^^^ ) ^ File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/simulator.py", line 548, in simulate_protocol_time_course self.model.update_parameters(cast(dict, pars.to_dict())) ~~~~~~~~~~~~~~~~~~~~~~~~~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^ File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/model.py", line 784, in update_parameters self.update_parameter(k, v) ~~~~~~~~~~~~~~~~~~~~~^^^^^^ File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/model.py", line 207, in wrapper return method(*args, **kwargs) File "/home/runner/work/MxlPy/MxlPy/src/mxlpy/model.py", line 749, in update_parameter raise KeyError(msg) KeyError: "Parameter 'k1' not found. Available parameters: ['k2', 'k_in', 'k_out']" """ The above exception was the direct cause of the following exception: KeyError Traceback (most recent call last) Cell In[5], line 1 ----> 1 tc = mc.protocol_time_course( 2 get_linear_chain_2v(), 3 time_points=np.linspace(0, 6, 21), 4 protocol=make_protocol( File ~/work/MxlPy/MxlPy/src/mxlpy/mc.py:427, in protocol_time_course(model, protocol, time_points, mc_to_scan, y0, max_workers, cache, worker, integrator) 424 if y0 is not None: 425 model.update_variables(y0) --> 427 res = parallelise( 428 partial( 429 _update_parameters_and_initial_conditions, 430 fn=partial( 431 worker, 432 protocol=protocol, 433 time_points=time_points, 434 integrator=integrator, 435 y0=None, 436 ), 437 model=model, 438 ), 439 inputs=list(mc_to_scan.iterrows()), 440 max_workers=max_workers, 441 cache=cache, 442 ) 443 return ProtocolScan( 444 to_scan=mc_to_scan, 445 protocol=protocol, 446 raw_results=dict(res), 447 ) File ~/work/MxlPy/MxlPy/src/mxlpy/parallel.py:173, in parallelise(fn, inputs, cache, parallel, max_workers, timeout, disable_tqdm, tqdm_desc) 171 for future in futures: 172 try: --> 173 key, value = future.result(timeout=timeout) 174 pbar.update(1) 175 results.append((key, value)) File ~/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/concurrent/futures/_base.py:456, in Future.result(self, timeout) 454 raise CancelledError() 455 elif self._state == FINISHED: --> 456 return self.__get_result() 457 else: 458 raise TimeoutError() File ~/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/concurrent/futures/_base.py:401, in Future.__get_result(self) 399 if self._exception is not None: 400 try: --> 401 raise self._exception 402 finally: 403 # Break a reference cycle with the exception in self._exception 404 self = None KeyError: "Parameter 'k1' not found. Available parameters: ['k2', 'k_in', 'k_out']"
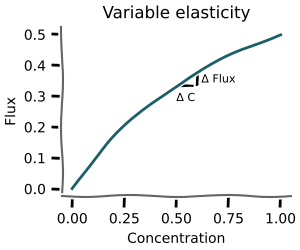
Metabolic control analysis¶
mxlpy further has routines for monte-carlo distributed metabolic control analysis.
This allows quantifying, whether the coefficients obtained from the MCA analysis are robust against parameter changes or whether they are just an artifact of a particular choice of parameters.
mc_elas = mc.variable_elasticities(
get_upper_glycolysis(),
variables={
"GLC": 0.3,
"G6P": 0.4,
"F6P": 0.5,
"FBP": 0.6,
"ATP": 0.4,
"ADP": 0.6,
},
to_scan=["GLC", "F6P"],
mc_to_scan=sample(
{
# "k1": LogNormal(mean=np.log(0.25), sigma=1.0),
# "k2": LogNormal(mean=np.log(1.0), sigma=1.0),
"k3": LogNormal(mean=np.log(1.0), sigma=1.0),
# "k4": LogNormal(mean=np.log(1.0), sigma=1.0),
# "k5": LogNormal(mean=np.log(1.0), sigma=1.0),
# "k6": LogNormal(mean=np.log(1.0), sigma=1.0),
# "k7": LogNormal(mean=np.log(2.5), sigma=1.0),
},
n=5,
),
)
_ = plot.violins_from_2d_idx(mc_elas)
plt.show()
0%| | 0/5 [00:00<?, ?it/s]
20%|██ | 1/5 [00:04<00:19, 4.97s/it]
40%|████ | 2/5 [00:05<00:06, 2.11s/it]
100%|██████████| 5/5 [00:05<00:00, 1.61it/s]
100%|██████████| 5/5 [00:05<00:00, 1.03s/it]
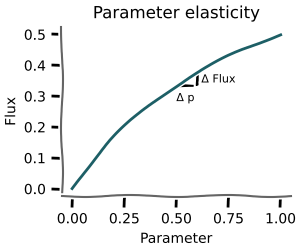
Parameter elasticities¶
 +
+
 =
=

elas = mc.parameter_elasticities(
get_upper_glycolysis(),
variables={
"GLC": 0.3,
"G6P": 0.4,
"F6P": 0.5,
"FBP": 0.6,
"ATP": 0.4,
"ADP": 0.6,
},
to_scan=["k1", "k2", "k3"],
mc_to_scan=sample(
{
"k3": LogNormal(mean=np.log(0.25), sigma=1.0),
},
n=5,
),
)
_ = plot.violins_from_2d_idx(elas)
plt.show()
0%| | 0/5 [00:00<?, ?it/s]
20%|██ | 1/5 [00:04<00:19, 5.00s/it]
40%|████ | 2/5 [00:05<00:06, 2.14s/it]
100%|██████████| 5/5 [00:05<00:00, 1.05s/it]
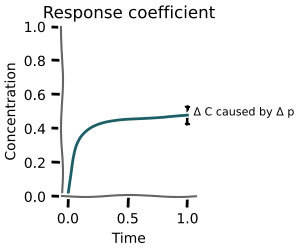
Response coefficients¶
 +
+
 =
=

# Compare with "normal" control coefficients
rc = mca.response_coefficients(
get_lin_chain_two_circles(),
to_scan=["vmax_1", "vmax_2", "vmax_3", "vmax_5", "vmax_6"],
)
_ = plot.heatmap(rc.variables)
mrc = mc.response_coefficients(
get_lin_chain_two_circles(),
to_scan=["vmax_1", "vmax_2", "vmax_3", "vmax_5", "vmax_6"],
mc_to_scan=sample(
{
"k0": LogNormal(np.log(1.0), 1.0),
"k4": LogNormal(np.log(0.5), 1.0),
},
n=10,
),
)
_ = plot.violins_from_2d_idx(mrc.variables, n_cols=len(mrc.variables.columns))
0%| | 0/5 [00:00<?, ?it/s]
20%|██ | 1/5 [00:05<00:20, 5.03s/it]
100%|██████████| 5/5 [00:05<00:00, 1.25it/s]
100%|██████████| 5/5 [00:05<00:00, 1.06s/it]
0%| | 0/10 [00:00<?, ?it/s]
0%| | 0/5 [00:00<?, ?it/s] 0%| | 0/5 [00:00<?, ?it/s] 0%| | 0/5 [00:00<?, ?it/s] 0%| | 0/5 [00:00<?, ?it/s] 40%|████ | 2/5 [00:00<00:00, 11.87it/s]
40%|████ | 2/5 [00:00<00:00, 10.26it/s] 40%|████ | 2/5 [00:00<00:00, 12.49it/s] 80%|████████ | 4/5 [00:00<00:00, 12.34it/s] 100%|██████████| 5/5 [00:00<00:00, 12.28it/s] 10%|█ | 1/10 [00:05<00:48, 5.44s/it]
80%|████████ | 4/5 [00:00<00:00, 10.45it/s] 80%|████████ | 4/5 [00:00<00:00, 12.41it/s]
100%|██████████| 5/5 [00:00<00:00, 10.83it/s] 100%|██████████| 5/5 [00:00<00:00, 12.45it/s] 0%| | 0/5 [00:00<?, ?it/s]
0%| | 0/5 [00:00<?, ?it/s] 0%| | 0/5 [00:00<?, ?it/s]
40%|████ | 2/5 [00:00<00:00, 13.43it/s] 40%|████ | 2/5 [00:00<00:00, 10.25it/s]
80%|████████ | 4/5 [00:00<00:00, 11.99it/s]
100%|██████████| 5/5 [00:00<00:00, 11.94it/s] 0%| | 0/5 [00:00<?, ?it/s] 80%|████████ | 4/5 [00:00<00:00, 10.27it/s]
40%|████ | 2/5 [00:00<00:00, 12.40it/s] 100%|██████████| 5/5 [00:00<00:00, 10.26it/s] 0%| | 0/5 [00:00<?, ?it/s]
80%|████████ | 4/5 [00:00<00:00, 12.26it/s] 40%|████ | 2/5 [00:00<00:00, 10.86it/s]
100%|██████████| 5/5 [00:00<00:00, 12.25it/s] 0%| | 0/5 [00:00<?, ?it/s]
80%|████████ | 4/5 [00:00<00:00, 10.85it/s] 40%|████ | 2/5 [00:00<00:00, 11.92it/s]
100%|██████████| 5/5 [00:00<00:00, 10.88it/s]
80%|████████ | 4/5 [00:00<00:00, 12.11it/s] 100%|██████████| 5/5 [00:00<00:00, 12.13it/s]
20%|██ | 1/5 [00:04<00:16, 4.11s/it]
20%|██ | 1/5 [00:03<00:15, 3.95s/it]
40%|████ | 2/5 [00:07<00:11, 3.69s/it]
40%|████ | 2/5 [00:07<00:10, 3.63s/it]
60%|██████ | 3/5 [00:10<00:07, 3.55s/it]
60%|██████ | 3/5 [00:10<00:07, 3.53s/it]
80%|████████ | 4/5 [00:14<00:03, 3.49s/it]
80%|████████ | 4/5 [00:14<00:03, 3.48s/it]
100%|██████████| 5/5 [00:17<00:00, 3.46s/it] 100%|██████████| 5/5 [00:17<00:00, 3.54s/it] 40%|████ | 4/10 [00:22<00:34, 5.74s/it]
100%|██████████| 5/5 [00:17<00:00, 3.45s/it] 100%|██████████| 5/5 [00:17<00:00, 3.51s/it] 60%|██████ | 6/10 [00:23<00:13, 3.33s/it]
100%|██████████| 10/10 [00:23<00:00, 2.33s/it]
First finish line
With that you now know most of what you will need from a day-to-day basis about monte carlo methods in mxlpy.Congratulations!
Advanced topics¶
Parameter scans¶
Vary both monte carlo parameters as well as systematically scan for other parameters
mcss = mc.scan_steady_state(
get_linear_chain_2v(),
to_scan=pd.DataFrame({"k1": np.linspace(0, 1, 3)}),
mc_to_scan=sample(
{
"k2": Uniform(1.0, 1.3),
"k3": LogNormal(mean=1.0, sigma=0.2),
},
n=10,
),
)
plot.violins_from_2d_idx(mcss.variables)
plt.show()
0%| | 0/10 [00:00<?, ?it/s]
0%| | 0/3 [00:00<?, ?it/s] 0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 165.72it/s] 10%|█ | 1/10 [00:04<00:44, 5.00s/it]
0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 151.86it/s] 0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 173.57it/s] 100%|██████████| 3/3 [00:00<00:00, 211.34it/s]
0%| | 0/3 [00:00<?, ?it/s] 0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 269.41it/s]
0%| | 0/3 [00:00<?, ?it/s] 50%|█████ | 5/10 [00:05<00:03, 1.27it/s]
0%| | 0/3 [00:00<?, ?it/s] 0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 204.04it/s] 0%| | 0/3 [00:00<?, ?it/s] 100%|██████████| 3/3 [00:00<00:00, 208.80it/s] 100%|██████████| 3/3 [00:00<00:00, 238.98it/s] 100%|██████████| 3/3 [00:00<00:00, 172.42it/s] 100%|██████████| 3/3 [00:00<00:00, 185.75it/s] 100%|██████████| 10/10 [00:05<00:00, 1.91it/s]
INFO:matplotlib.category:Using categorical units to plot a list of strings that are all parsable as floats or dates. If these strings should be plotted as numbers, cast to the appropriate data type before plotting.
INFO:matplotlib.category:Using categorical units to plot a list of strings that are all parsable as floats or dates. If these strings should be plotted as numbers, cast to the appropriate data type before plotting.
INFO:matplotlib.category:Using categorical units to plot a list of strings that are all parsable as floats or dates. If these strings should be plotted as numbers, cast to the appropriate data type before plotting.
INFO:matplotlib.category:Using categorical units to plot a list of strings that are all parsable as floats or dates. If these strings should be plotted as numbers, cast to the appropriate data type before plotting.
# FIXME: no idea how to plot this yet. Ridge plots?
# Maybe it's just a bit much :D
mcss = mc.scan_steady_state(
get_linear_chain_2v(),
to_scan=mb2.cartesian_product(
{
"k1": np.linspace(0, 1, 3),
"k2": np.linspace(0, 1, 3),
}
),
mc_to_scan=sample(
{
"k3": LogNormal(mean=1.0, sigma=0.2),
},
n=10,
),
)
mcss.variables.head()
0%| | 0/10 [00:00<?, ?it/s]
0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s]
11%|█ | 1/9 [00:00<00:06, 1.33it/s] 11%|█ | 1/9 [00:00<00:06, 1.32it/s] 11%|█ | 1/9 [00:00<00:06, 1.29it/s] 11%|█ | 1/9 [00:00<00:06, 1.28it/s]
44%|████▍ | 4/9 [00:01<00:01, 2.88it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.87it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.80it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.80it/s]
78%|███████▊ | 7/9 [00:02<00:00, 3.36it/s] 78%|███████▊ | 7/9 [00:02<00:00, 3.35it/s] 100%|██████████| 9/9 [00:02<00:00, 3.94it/s] 100%|██████████| 9/9 [00:02<00:00, 3.93it/s] 78%|███████▊ | 7/9 [00:02<00:00, 3.29it/s] 100%|██████████| 9/9 [00:02<00:00, 3.85it/s] 10%|█ | 1/10 [00:07<01:06, 7.34s/it]
78%|███████▊ | 7/9 [00:02<00:00, 3.31it/s] 100%|██████████| 9/9 [00:02<00:00, 3.87it/s]
0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s] 0%| | 0/9 [00:00<?, ?it/s]
11%|█ | 1/9 [00:00<00:06, 1.30it/s] 11%|█ | 1/9 [00:00<00:06, 1.27it/s] 11%|█ | 1/9 [00:00<00:05, 1.36it/s] 11%|█ | 1/9 [00:00<00:06, 1.26it/s]
44%|████▍ | 4/9 [00:01<00:01, 2.81it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.86it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.77it/s] 44%|████▍ | 4/9 [00:01<00:01, 2.78it/s]
78%|███████▊ | 7/9 [00:02<00:00, 3.29it/s] 100%|██████████| 9/9 [00:02<00:00, 3.86it/s] 0%| | 0/9 [00:00<?, ?it/s] 50%|█████ | 5/10 [00:09<00:08, 1.62s/it]
78%|███████▊ | 7/9 [00:02<00:00, 3.32it/s] 78%|███████▊ | 7/9 [00:02<00:00, 3.27it/s] 100%|██████████| 9/9 [00:02<00:00, 3.91it/s] 100%|██████████| 9/9 [00:02<00:00, 3.82it/s] 0%| | 0/9 [00:00<?, ?it/s] 78%|███████▊ | 7/9 [00:02<00:00, 3.27it/s] 100%|██████████| 9/9 [00:02<00:00, 3.82it/s]
11%|█ | 1/9 [00:00<00:03, 2.38it/s] 11%|█ | 1/9 [00:00<00:03, 2.31it/s]
44%|████▍ | 4/9 [00:00<00:00, 5.32it/s] 44%|████▍ | 4/9 [00:00<00:00, 5.25it/s]
78%|███████▊ | 7/9 [00:01<00:00, 6.26it/s] 100%|██████████| 9/9 [00:01<00:00, 7.30it/s] 90%|█████████ | 9/10 [00:11<00:00, 1.11it/s]
78%|███████▊ | 7/9 [00:01<00:00, 6.21it/s] 100%|██████████| 9/9 [00:01<00:00, 7.22it/s] 100%|██████████| 10/10 [00:11<00:00, 1.11s/it]
| x | y | |||
|---|---|---|---|---|
| k1 | k2 | |||
| 0 | 0.0 | 0.0 | NaN | NaN |
| 0.5 | 2.0 | 1.0 | ||
| 1.0 | 1.0 | 1.0 | ||
| 0.5 | 0.0 | NaN | NaN | |
| 0.5 | 2.0 | 1.0 |
Custom distributions¶
If you want to create custom distributions, all you need to do is to create a class that follows the Distribution protocol, e.g. implements a sample function.
For API consistency, the sample method has to take rng argument, which can be ignored if not applicable.
from dataclasses import dataclass
from typing import TYPE_CHECKING
if TYPE_CHECKING:
from mxlpy.types import Array
@dataclass
class MyOwnDistribution:
loc: float = 0.0
scale: float = 1.0
def sample(
self,
num: int,
rng: np.random.Generator | None = None,
) -> Array:
if rng is None:
rng = np.random.default_rng()
return rng.normal(loc=self.loc, scale=self.scale, size=num)
sample({"p1": MyOwnDistribution()}, n=5)
| p1 | |
|---|---|
| 0 | 0.834082 |
| 1 | -0.146356 |
| 2 | -0.080290 |
| 3 | -2.393945 |
| 4 | -0.236137 |